How artificial intelligence is helping scientists find a coronavirus treatment
More than 50,000 academic articles have been written about COVID-19 since the virus appeared in November.
The volume of new information isn't necessarily a good thing.
Not all of the recent coronavirus literature has been peer reviewed, while the sheer number of articles makes it challenging for accurate and promising research to stand out or be further studied.
Computer science and linguistics professor James Pustejovsky is leading a Brandeis team in creating an artificial intelligence platform called Semantic Visualization of Scientific Data—or SemViz—that can sort through the growing mass of published work on coronavirus and help biologists who study the disease gain insights and notice patterns and trends across research that could lead to a treatment or cure.
Pustejovsky, an expert in theoretical and computational modeling and language, is partnering with colleagues at Tufts University, Harvard University, the University of Illinois, and Vassar College. He discussed his work with BrandeisNOW.
Can you provide a bird's-eye view of the way you've applied your background as a computational linguist to current coronavirus research?
I'm a researcher who focuses on language and extracting information from large amounts of text, like the COVID-19 dataset, which now includes more than 50,000 academic articles. Biologists on the front lines of coronavirus are trying to find connections between genes, proteins and drugs, and how they interact with the virus in the cells of the human body.
SemViz combs through the existing papers and manuscripts and enables scientists to make connections and generalizations that are not obvious from reading one paper at a time.
So how might a biologist studying coronavirus actually use SemViz?
This tool gives a rapid way for biologists studying coronavirus to see a global overview of inhibitors, regulators, and activators of genes and proteins involved in the disease.
For example, what are the drugs and proteins regulating the receptor for the COVID-19 virus? This could help discover therapies that decrease the expression of the receptor for the virus in patients' lungs. This is important because millions of people currently take blood pressure medicines that can alter this receptor and possibly increase their risk of contracting the disease.
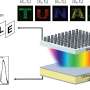
SemViz creates a visualization landscape that helps biologists make both global and specific connections between human genes, drugs, proteins and viruses. The overall program I'm working on contains three components: two semantic visualization outputs based on the entire coronavirus research dataset, as well as a natural language-based question-answering application.
What's the language application grid and how does it work?
It is essentially a computer-based "reading machine" that interprets tens of thousands of research articles on coronavirus and presents the results of this process to biologists in a form that is visually accessible and easily analyzed and interpreted.
It is more informative than a search engine, because it utilizes a host of language understanding tools and AI that can be applied to different domains (economics, news, science, literature) and text types (tweets, articles, books, email).
What are the implications of SemViz?
I think it's hard to overstate the challenge brought about by information overload, particularly now with the coronavirus literature.
Biologists are interested in the mechanisms and functions of specific chemicals and proteins. SemViz can be the roadmap that scientists use to sort through large amounts of research to find these kinds of functions and relationships.