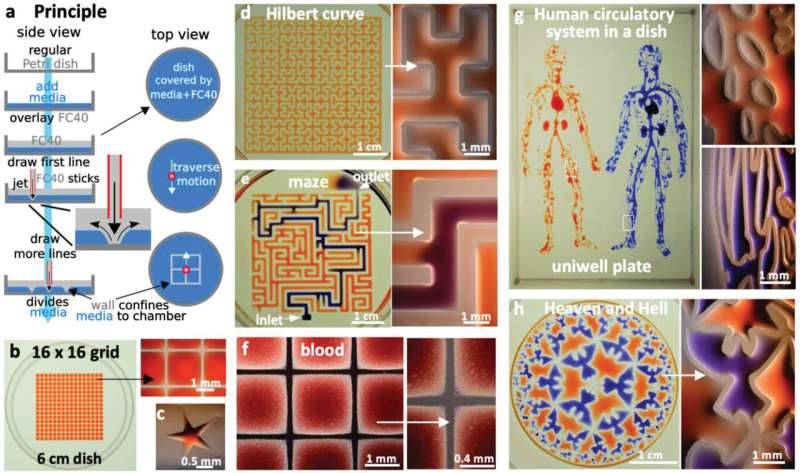
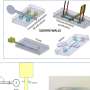
Principle. a) A “microjet” of an immiscible fluorocarbon (FC40) is projected through FC40 and media on to the bottom of a dish; media is pushed aside, and FC40 sticks to the bottom (it wets polystyrene better than media). Moving the jet now draws lines to form a grid with FC40 walls. b) A 16 × 16 grid (interchamber spacing 1.9 mm; 600 nL red dye added to each chamber; zoom shows fluid walls). c) Star‐shaped walls (red dye added after jetting). d) A square with one internal wall shaped like a Hilbert curve (a continuous line with many 90° and 45° turns). Red dye was added at many points; it diffused around the wall throughout the square. Zoom shows some turns. e) Path through a maze with a single inlet and outlet. The circuit was built by jetting FC40 through media plus red dye; when blue dye is pumped into the inlet, it takes the path of least resistance through the maze. Zoom shows some walls. f) Part of 16 × 16 grid where FC40 is jetted through blood instead of media. Zoom shows the jet leaves no cells in what are now FC40 walls. g) Human circulatory system in a uniwell plate with dimensions of a 96‐well plate. Red and blue dyes were infused into major “arteries” and “veins” which then flow/diffuse throughout each system. Zooms show regions in white rectangles. h) Walls were built in design inspired by “Circle Limit IV (Heaven and Hell)” by M.C. Escher – WikiArt. Patterns for Figure 1d,e,g were obtained from the following sources: © Wikimedia Commons (author Braindrain0000), Can Stock Photo Inc. (author Taui), pngegg.com, respectively. Images used to inspire Figure 1d and 1e were obtained and used with permission from Wikimedia Commons (name "Hilbert curve.png", author Zbigniew Fiedorowicz) and Can Stock Photo Inc. (image number csp49511651, author Taui)”. Image used to inspire Figure 1g is a public domain image. Image used to inspire Figure 1h was from Sazdanovic et al. and is used under the terms of the CC‐BY license, Copyright 2012, Radmila Sazdanovic, published by MDPI. Credit: Advanced Science, doi: 10.1002/advs.202001854
Biomedical applications in life sciences can greatly benefit from microfluidics devices; however, the technology is suboptimal for rapid production and applications in biolabs. For instance, solid opaque walls of conventional microfluidic devices prevent biologists from providing adequate physical and optical access to their biological samples. Therefore, there is a growing need to engineer optimized microfluidics for efficient workflow. In a new study, Cristian Soitu and a research team at the Walsh and Cook Research Groups in the Department of Engineering Science and the Sir William Dunn School of Pathology at the University of Oxford, U.K., described a new contactless, microfluidics circuit fabrication method.
The team used standard petri dishes as the platform on which they developed the circuits using commonly available biolab tools, including a syringe pump, dispensing needles, cell culture media and an immiscible fluorocarbon (FC40) compound (a material freely permeable to oxygen and carbon dioxide). During the experiments, they "jetted" the FC40 from a dispensing needle through the medium present in the petri dish to reproducibly manufacture complex and highly accurate microfluidics structures in minutes. The technique allows the miniaturization of some common workflows in biology coupled with a flexible and diverse approach, well suited for widespread adoption in biomedicine. The work is now published on Advanced Science.
The new technique—jet-printing microfluidic devices on demand
The primary author of the study; bioengineer and Ph.D. researcher Cristian Soitu described the novelty of their frugal technique as "a contactless method to build fluid walls using materials commonly found in most biolabs." While detailing the speed and efficacy of their method, he added that "the fluid walls can be quickly built into any imaginable 2-D (two dimensional) circuit and we fabricated a human circulatory system in a microplate under 30 minutes, as a proof of concept." The new study is based on preceding work by the same research group who established 'freestyle fluidics' to use fluids as building blocks and confine liquids with fluid walls instead of solid walls.
Path through a maze with a single inlet and outlet. The circuit was built by jetting FC40 through media plus red dye; when blue dye is pumped into the inlet, it takes the path of least resistance through the maze. Credit: Advanced Science, doi: 10.1002/advs.202001854.
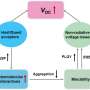
A key feature of the new approach is that it can overcome the Poisson limit during mammalian cell culture cloning. The Poisson statistics dictate that most cells cultured across wells in a microplate will be discarded after cell splitting and cloning protocols, thereby wasting cells and reagents. The jetting method will allow researchers to beat the Poisson limit by building isolated wells (known as Voronoi regions) to subculture the mammalian cells steadily in fresh media for a week, to harvest cell clones thereafter. Conventional open microfluidic devices too can be integrated into biomedical workflows, but they require substrate etching, surface treatment, contact or a combination of efforts to confine fluids and biological structures. Such complex manufacturing processes can deter biologists who favor efficient and flexible prototyping without compromising biocompatibility. As a result, the described jetting technique can provide a new hands-on approach to overcome existing limits of microfluidics.
The experimental approach
During the experiments, the researchers covered the bottom of a standard tissue-culture dish with a film of cell growth media and an overlay of FC40 to prevent evaporation. By moving the microjet sideways, they created a grid with 256 chambers in under two minutes on the FC40 layer formed on the petri dish. In this contactless method, the nozzle did not touch the dish or media, preventing cross-contamination. Conventional flow systems are highly specialized and built in, and are therefore difficult to integrate into existing workflows. Comparatively, the perfusion circuit jetted within a standard petri dish here, allowed perfusion within 15 minutes to feed myoblasts (embryonic precursors of muscle cells) to subsequently generate myotubes (muscle fibers formed via myoblast fusion) within a week. Soitu et al. additionally highlight the ability to sub-culture cells without using trypsin/EDTA (conventional reagents), given that a whole or part of the colonies will be dislodged by the FC40 jetting process to transfer to new chambers. The method therefore allowed them to sample a single colony several times as it grew.
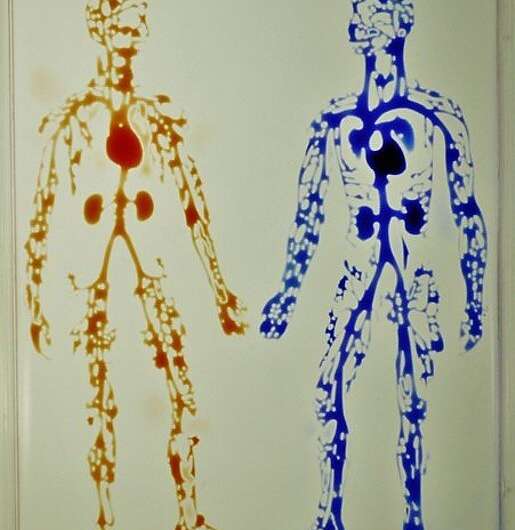
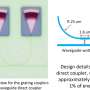
Human circulatory system in a uniwell plate with dimensions of a 96‐well plate. Red and blue dyes were infused into major “arteries” and “veins” which then flow/diffuse throughout each system. Credit: Advanced Science, doi: 10.1002/advs.202001854
Reproducibility of the method
To demonstrate the reproducibility of the method, the team used jetting to build walls in the form of a star-shaped chamber, as a square enclosing a single continuous wall shaped like a Hilbert curve and a microfluidic maze with blue dye inserted to solve a maze and find the exit. The method is not restricted to the prescribed media and can be replaced with whole human blood for specific experiments. The patterns themselves include the "the human circulatory system" and the "circle limit IV" (Heaven and Hell) by M.C. Escher (WikiArt), which are diverse and flexible. The efforts highlight the potential to reproduce almost any pre-existing pattern. The resulting fluid walls were sufficiently stable; therefore, the researchers could carry the constructs between labs, incubators and microscopes, much like any dish filled with liquid.
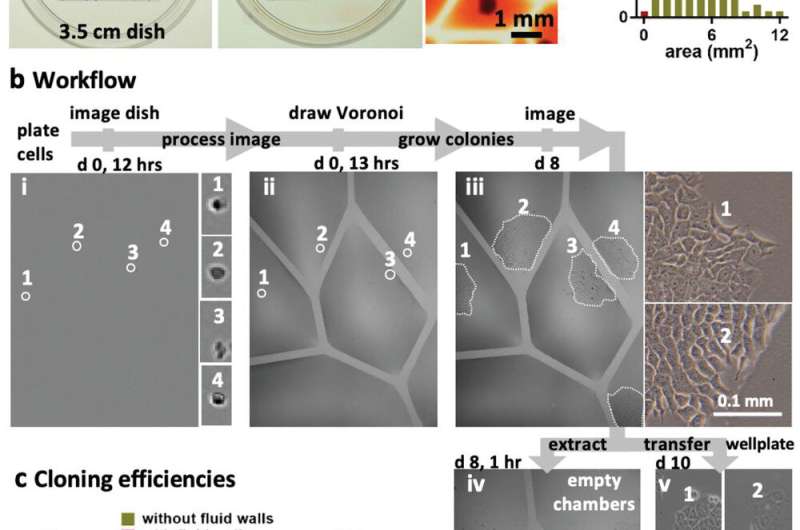
Beating the Poisson limit during cell cloning. a) Principle illustrated using 100 randomly distributed dots in a 2 cm square printed on clear film placed under a 35 mm dish. i) Dish + dots. ii) After recording dot positions, the Voronoi diagram is determined, polygonal walls built around each dot, and each polygonal chamber filled with dye (zoom shows some chambers). iii) Distribution of areas in this Voronoi diagram. Two polygons had area (A) <1 mm2, a threshold determined by the minimum area (hence volume) the infusing needle can access. b) Workflow illustrated using NM18 cells (12 cells cm−2 or ≈48 in 2 × 2 cm square) plated in a 35 mm dish. Phase‐contrast images are of the same region, and numbered zooms show magnifications of different founder cells and their colonies. i) Cells 1–4 12 h after plating. ii) The same cells after jetting surrounding polygons. iii) By day 8, cells develop into colonies (outlined by dotted lines). iv) Emptied polygons (achieved by adding/removing PBS, adding trypsin, incubation, and removing most cells). Overemptying some chambers leaves FC40 (dark blobs) attached to the bottom. v) After removal, each clone was reseeded in a well in a 12‐well plate, and imaged 2 days later. c) Cloning efficiencies. Each cell type was seeded (12 cells cm−2) in 10 × 35 mm dishes, colonies counted after 8 days, and cloning efficiencies calculated. Five dishes were used to assess cloning efficiencies conventionally (“without fluid walls”), and five using polygons (“with fluid walls”). There was no or little significant difference between the two approaches (unpaired t test for HEK, ES, and NM18 gave p = 0.488, 0.3, and 0.033, respectively). Credit: Advanced Science, doi: 10.1002/advs.202001854
Building a complex perfusion circuit with constant flow
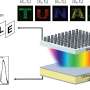
The team then engineered a complex circuit via jet printing to provide a steady flow of fresh media for seven days to maintain cell growth in an array of chambers. To accomplish this, they designed a circuit where each chamber received fresh media and demonstrated circuit functionality using mouse myoblasts that differentiated into mature myotubes within seven days. If required, the team can disconnect any chamber from the adjoining circuit by building new FC40 walls.
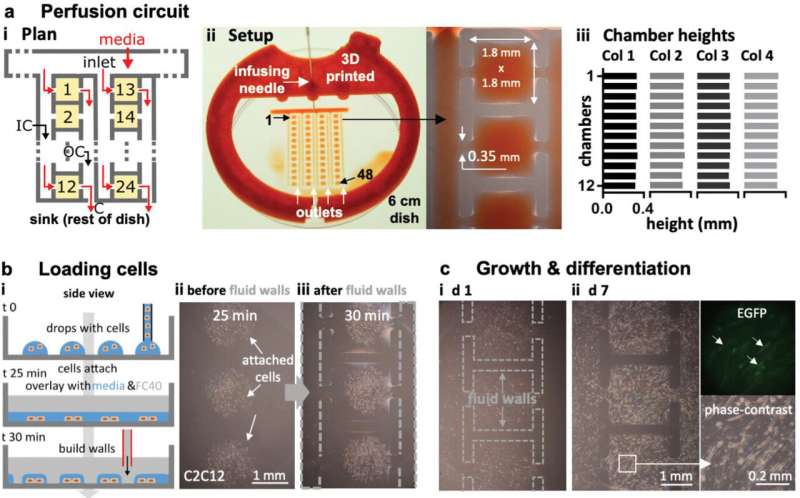
Perfusion circuit for continuous feeding 48 sets of differentiating myoblasts. a) Plan of the first two columns in the four‐column circuit i). Media (±dye) driven by an external pump into the inlet flows (red arrows) through an input conduit (IC), one cell chamber (chambers 1, 2, …, 24 are orange), an output conduit (OC), a choke (C), and out into the dish (the sink). Gray lines indicate FC40 walls. ii) Setup. Red dye is pumped into the inlet through a 3D‐printed adapter (red) that fits on a 6 cm dish, through 48 chambers, and out into the dish (zoom shows some chambers). iii) All chambers have similar maximum heights (average ± SD = 325 ± 6.3 µm; assessed using fluorescent dyes) showing that all experience similar pressures, shear stresses, and flows. b) Growth and differentiation of mouse C2C12 myoblasts. i) Workflow. Myoblasts are deposited in drops, the dish filled with media and FC40 added, and finally the circuit is jetted around them. ii,iii) Cells before/after building fluid walls. Dashed lines: edges of some walls. c) Images of chambers as media replenished (1 µL per day per chamber); myoblasts grow to fill chambers, form syncytia, and express EGFP‐DOK7 (day 7). Fluorescence and phase‐contrast zooms: arrows mark fluorescing syncytia each containing >20 nuclei with length >200 µm—indicative of differentiation into myotubes. The speed and extent of growth and differentiation is like that of cells grown conventionally. Credit: Advanced Science, doi: 10.1002/advs.202001854
In this way, Cristian Soitu and colleagues developed a rapid and precise technique to generate microfluidic circuits on a standard polystyrene petri dish. They projected a microjet of PC40 to the bottom of the dish to build a liquid wall pinned by interfacial forces. The team proved the ability to (1) build stable fluid circuits with any imaginable 2-D shape within seconds to minutes, (2) drive flow via conduits without using external pumps, and (3) create valves that can be opened and closed at will. Key features of the approach will allow biologists to beat the Poisson limit, generate a robust perfusion system and subculture attached cells without conventional reagents (such as trypsin/EDTA). The research team will optimize the device and overcome existing limits to construct ever-smaller fluid circuits capable of retaining miniaturized volumes of media. The diversity and flexibility of this approach is expected to lead to widespread adoption of the technique for a variety of applications in biomedicine.
More information: Cristian Soitu et al. Jet‐Printing Microfluidic Devices on Demand, Advanced Science (2020). DOI: 10.1002/advs.202001854
Eric K. Sackmann et al. The present and future role of microfluidics in biomedical research, Nature (2014). DOI: 10.1038/nature13118
Cristian Soitu et al. Microfluidic chambers using fluid walls for cell biology, Proceedings of the National Academy of Sciences (2018). DOI: 10.1073/pnas.1805449115
Journal information: Nature , Proceedings of the National Academy of Sciences
© 2020 Science X Network