Credit: Kruiper et al.
Scientific texts, such as research articles or reviews, can sometimes be difficult to analyze and understand, particularly for non-expert readers. In recent years, engineers have thus tried to develop approaches that can automatically extract the most important information from dense scientific texts, which can then be used to guide readers and aid their understanding of the texts.
Some of the information extraction (IE) systems developed so far, however, can only extract a fraction of a text's content, while others have been found to perform poorly on texts that contain long and complex sentences. In a recent paper pre-published on arXiv, researchers at Heriot-Watt University in Scotland introduced a new IE approach that combines two of the most commonly used techniques for extracting information from scientific texts.
"Our research at Heriot-Watt University aims to support nature-inspired problem solving," Ruben Kruiper, one of the researchers who carried out the study, told TechXplore. "The idea is that engineers need help finding relevant information in biology research papers. A major problem is that engineers and the industry in general lack the biological expertise to even recognise relevant information."
Computer scientists who are trying to understand biology papers and apply concepts presented in them in their research often struggle to understand biological jargon and quickly determine whether an article is worth reading in more depth. These issues are also often encountered by other readers who lack expertise in the scientific domain they are reading about.
"Sometimes, even experts spend hours trying to identify the central theme and concepts in newly published literature," Kruiper said. "In our work, we try to support all readers of scientific texts by providing a summary view of the central concepts discussed in them."
Typically, there are two kinds of systems to extract information from scientific texts: narrow and open IE systems. The first type works by precisely identifying a handful of relations between different notions contained in the text, for instance focusing on drug-gene interactions in pharmacological studies. For this type of system to work, however, researchers need to specify the type of relations that it should be looking for.
The second type of IE system implements a scattershot-type method, for instance unveiling pairs of nouns and phrases that are connected by a verb. A limitation of this method is that it gives researchers very little control over the facts they are extracting. Moreover, the complex syntax of sentences typically contained in scientific texts can affect the system's performance, resulting in the extraction of wrong, incomplete or irrelevant information.
Credit: Kruiper et al.
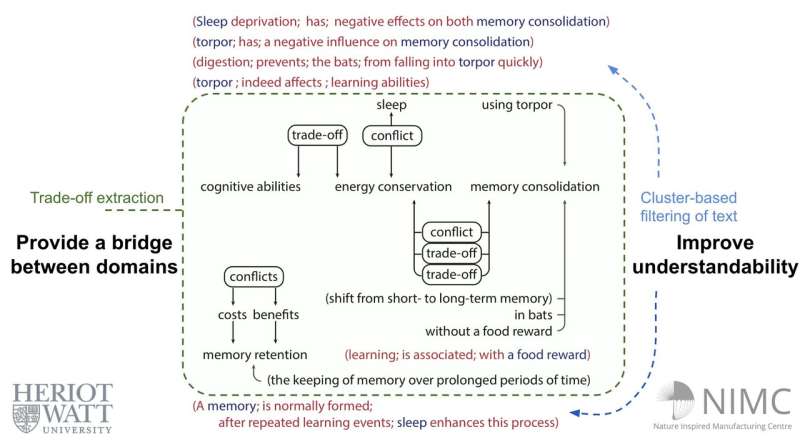
"Our approach combines the outputs of both types of systems, a task that we call semi-open relation extraction," Kruiper said. "We extract the information we want precisely, and then use these extractions to filter the results of a scattershot system."
The system developed by Kruiper and his colleagues finds a unique balance between the accuracy and flexibility of the two most commonly used IE techniques. The researchers ran it on a corpus of 10,000 biology-related texts and found that it achieved remarkable performance, successfully extracting the most crucial information contained in them.
"We showed that that our semi-open relation extraction approach is worthwhile," Kruiper said. "Filtering the facts extracted by a scattershot system improves the overall quality, while greatly reducing the overwhelming number of facts in a document. The combined approach we developed can identify such a central relation with reasonable accuracy, while also identifying closely related facts."
The semi-open relation extraction system introduced by this team of researchers can automatically extract the main points contained in a scientific article, allowing readers to quickly decide whether it's worth reading it more in depth and identify sections that might be of interest to them.
The IE system's code is publicly available online and can be accessed on Kruiper's GitHub page. In the future, it could prove useful for researchers or engineers who are looking for scientific information on a topic that is outside of their field of expertise or who need to browse through large amounts of research articles quickly.
So far, the researchers merely explored the feasibility of combining narrow and open IE systems. In their next studies, they would like to compile a dataset that could be used to train IE techniques, further pushing the boundaries of IE from scientific texts.
"There is much room for improving and simplifying the overall system," Kruiper said. "The current setup does, however, already enable the collection of a larger and more comprehensive dataset. Preparing such a dataset to train new systems, as well as using the current setup in Biomimetic case studies, will provide valuable insight in the types of information we want to be extracting precisely."
Kruiper and his colleagues work at Heriot-Watt University's Interaction Lab and Nature Inspired Manufacturing Centre (NIMC), which has the key mission of supporting companies in their search for more sustainable manufacturing solutions. In addition to conducting further research, therefore, they are currently seeking funding from the UK government and companies that could back their work and support them in developing new technology.
More information: In layman's terms: semi-open relation extraction from scientific texts. arXiv:2005.07751 [cs.CL]. arxiv.org/abs/2005.07751
© 2020 Science X Network