Researchers develop a self-supervised learning framework that leverages the large amounts of unlabeled data that other models can’t. Credit: Mechanical and AI Lab, Carnegie Mellon University
Predicting molecular properties quickly and accurately is important to advancing scientific discovery and application in areas ranging from materials science to pharmaceuticals. Because experiments and simulations to explore potential options are time-consuming and costly, scientists have investigated using machine learning (ML) methods to aid in computational chemistry research. But, most ML models can only make use of known, or labeled, data. This makes it nearly impossible to predict with accuracy the properties of novel compounds.
In an industry like drug discovery, there are millions of molecules from which to select for use in a potential drug candidate. A prediction error as small as 1% can lead to the misidentification of more than ten thousand molecules. Improving the accuracy of ML models with limited data will play a vital role in developing new treatments for disease.
While the amount of labeled molecule data is limited, there is a rapidly growing amount of feasible, but unlabeled, data. Researchers at Carnegie Mellon University's College of Engineering pondered if they could use this large volume of unlabeled molecules to build ML models that could perform better on property predictions than other models.
Their work culminated in the development of a self-supervised learning framework named MolCLR, short for Molecular Contrastive Learning of Representations with Graph Neural Networks (GNNs). The findings were published in the journal Nature Machine Intelligence.
"MolCLR significantly boosts the performance of ML models by leveraging approximately 10 million unlabeled molecule data," said Amir Barati Farimani, assistant professor of mechanical engineering.
For a simple explanation of labeled vs. unlabeled data, Ph.D. student Yuyang Wang suggested thinking of two sets of images of dogs and cats. In one set, each animal is labeled with the name of its species. In the other set, no labels accompany the images. To a human, the difference between the two types of animals might be obvious. But to a machine learning model, the difference isn't clear. The unlabeled data is therefore not reliably useful. Applying this analogy to the millions of unlabeled molecules that could take humans decades to manually identify, the critical need for smarter machine learning tools becomes obvious.
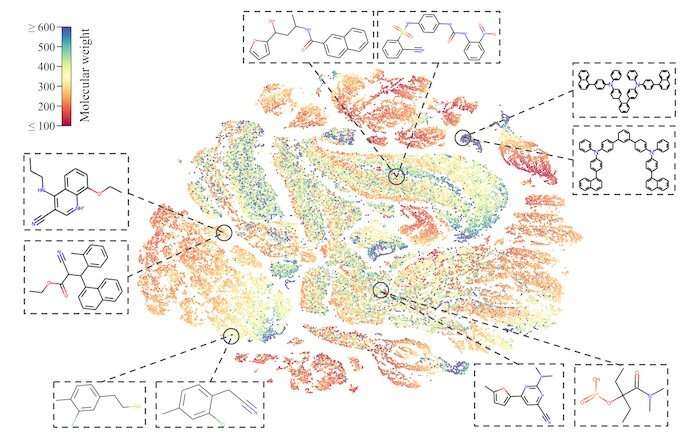
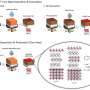
The research team sought to teach its MolCLR framework how to use unlabeled data by contrasting positive and negative pairs of augmented molecule graph representations. Graphs transformed from the same molecule are considered a positive pair, while those from different molecules are negative pairs. By this means, representations of similar molecules stay close to each other, while distinct ones are pushed far apart.
The researchers had applied three graph augmentations to remove small amounts of information from the unknown molecules: atom masking, bond deletion, and subgraph removal. In atom masking, a piece of information about a molecule is eliminated. In bond deletion, a chemical bond between atoms is erased. A combination of both augmentations results in subgraph removal. Through these three types of changes, the MolCLR was forced to learn intrinsic information and make correlations.
When the team applied MolCLR to ClinTox, a database used to predict drug toxicity, MolCLR significantly outperformed other ML baseline models. On another database, Tox21, MolCLR stood out from the other ML models with the potential to distinguish which environmental chemicals posed the most severe threats to human health.
"We've demonstrated that MolCLR bears promise for efficient molecule design," said Barati Farimani. "It can be applied to a wide variety of applications, including drug discovery, energy storage, and environmental protection."
More information: Yuyang Wang et al, Molecular contrastive learning of representations via graph neural networks, Nature Machine Intelligence (2022). DOI: 10.1038/s42256-022-00447-x
Journal information: Nature Machine Intelligence