August 27, 2018 feature
An evaluation of machine learning to identify bacteraemia in SIRS patients
A team of researchers at the Medical University of Vienna has recently evaluated the effectiveness of machine learning strategies to identify bacteraemia in patients affected by systemic inflammatory response syndrome (SIRS). Their study, published in Scientific Reports, gathered discouraging results, as machine learning methods could not achieve better accuracy than current diagnostic techniques.
Bacteraemia is a frequent medical condition characterized by the presence of bacteria in the blood, with a mortality rate ranging between 13 percent and 21 percent. Past research suggests that a number of factors are associated with the risk of developing this condition, including advanced age, urinary or indwelling vascular catheter, chemotherapy, and immunosuppressive therapies.
Diagnosing bacteraemia early is of crucial importance for the survival of affected patients, as they require prompt treatment with appropriate antibiotics. Currently, the analysis of blood culture (BC) is the main method for diagnosing the condition. However, this method is far from ideal, as it is often hard to determine who should undergo BC analysis, the results need around three days to be processed, and it can lead to around 8 percent of false positives.
Researchers are hence trying to identify biomarkers or prediction tools that could better identify patients having a high bacteraemia risk. So far, procalcitonin (PCT) has been found to be the best biomarker for detecting the condition, with a pooled sensitivity of 76 percent and a pooled specificity of 69 percent.
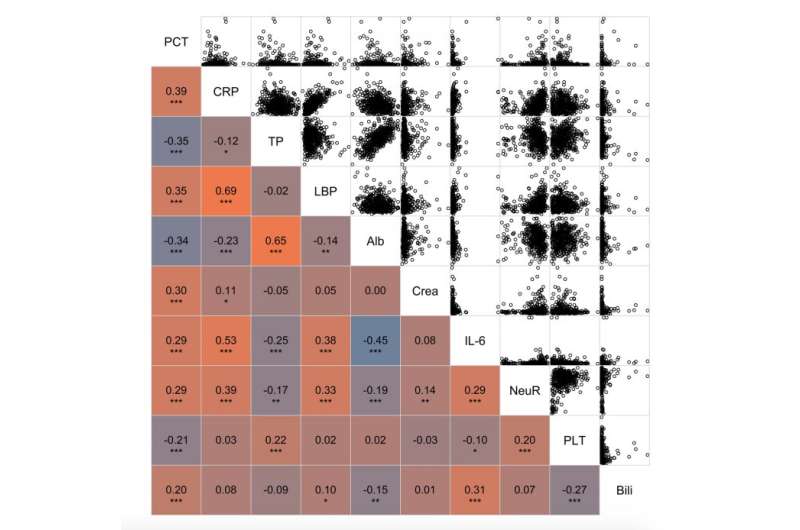
In their study, the researchers investigated whether machine learning strategies could improve the diagnostic performance of PCT in identifying bacteraemia, particularly in patients with two or more symptoms of SIRS who did not require BC analysis. They collected data from 466 patients who met the criteria and used a 29 parameter panel of clinical data, cytokine expression levels and standard lab markers to train their predictive model.
"The main objective of our study was to show whether the presence of bacteria in a patient's blood after they have exhibited inflammatory reactions can be predicted early on and better than currently possible, using laboratory parameters and machine learning," Georg Dorffner, one of the researchers who carried out the study, told Tech Xplore. "For that purpose, we conducted a large study with patients from our university clinic (AKH Vienna) to collect the necessary data."
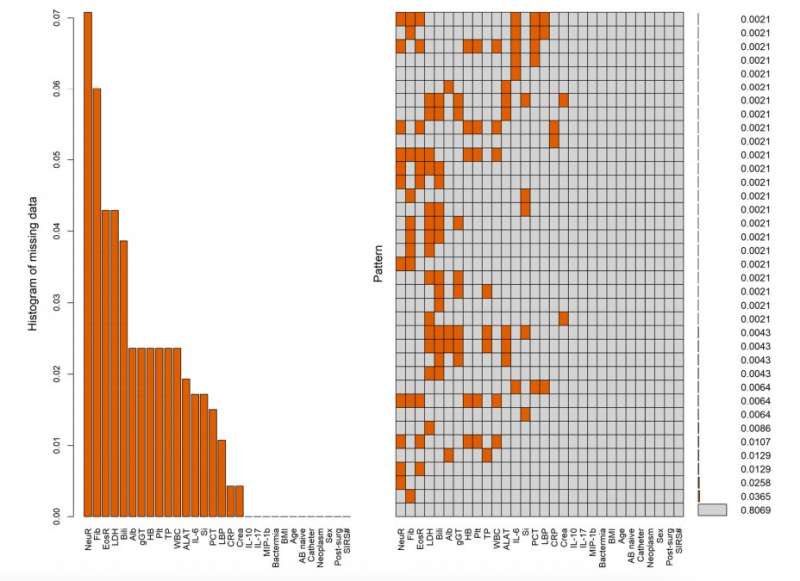
Doffner and his colleagues used a few predictive models that are popular within the field of machine learning, evaluating their respective effectiveness. They particularly focused on two models, one using neural networks and the other called random forest.
"One of the models we used is called 'neural network,' and finds good combinations of laboratory values such as to also make non-linear (i.e. non-proportional) predictions," Dorffner explained. "Another one—actually the best performing one—is called 'random forest,' and consists of a large number of so-called decision trees, where each tree tries to make a series of step-wise decisions, each based on a single laboratory value, as to what is the best prediction. These trees then all work together like a committee (hence, the name 'forest')."
In their study, the random forest strategy achieved the best results on predicting bacteraemia. However, it achieved a diagnostic accuracy equal to that of the biomarker PCT, suggesting that popular machine learning techniques are unable to predict the condition better than currently employed methods.
"Our most meaningful finding was that a set of several laboratory values could not lead to a better prediction than the one value that everyone else is using, namely the level of procalcitonin in the blood," Dorffner explained. "So machine learning did not really help advance the clinical routine in this case. It was still a worthwhile endeavor, as our results tell other researchers that the problem is not apparently predictable, sparing them unnecessary further work in this direction."
While the results collected by Dorffner and his colleagues were somewhat disappointing, they offer valuable insight for future research, outlining the difficulties of using machine learning to identify bacteraemia in SIRS patients.
"We are now focusing on other clinical applications where machine learning is likely more promising to advance predictions or diagnoses," Dorffner said. "For instance, together with cardiologists we are developing an MR-image-based learning system for detecting the rare but important disease of cardiac amyloidosis."
More information: Franz Ratzinger et al. Machine learning for fast identification of bacteraemia in SIRS patients treated on standard care wards: a cohort study, Scientific Reports (2018). DOI: 10.1038/s41598-018-30236-9
S.H. Hoeboer et al. The diagnostic accuracy of procalcitonin for bacteraemia: a systematic review and meta-analysis, Clinical Microbiology and Infection (2015). DOI: 10.1016/j.cmi.2014.12.026
© 2018 Medical Xpress