Recognizing disease using less data
As artificial intelligence systems learn to better recognize and classify images, they are becoming highly-reliable at diagnosing diseases, such as skin cancer, from medical images. But as good as they are at detecting patterns, AI won't be replacing your doctor any time soon. Even when used as a tool, image recognition systems still require an expert to label the data, and a lot of data at that: it needs images of both healthy patients and sick patients. The algorithm finds patterns in the training data and when it receives new data, it uses what it has learned to identify the new image.
One challenge is that it's time-consuming and costly for an expert to obtain and label each image. To address this issue, a group of researchers from Carnegie Mellon University's College of Engineering, including Professors Hae Young Noh and Asim Smailagic, teamed up to develop an active learning technique that uses a limited data set to achieve a high degree of accuracy in diagnosing diseases like diabetic retinopathy or skin cancer.
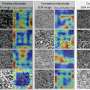
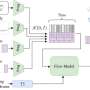
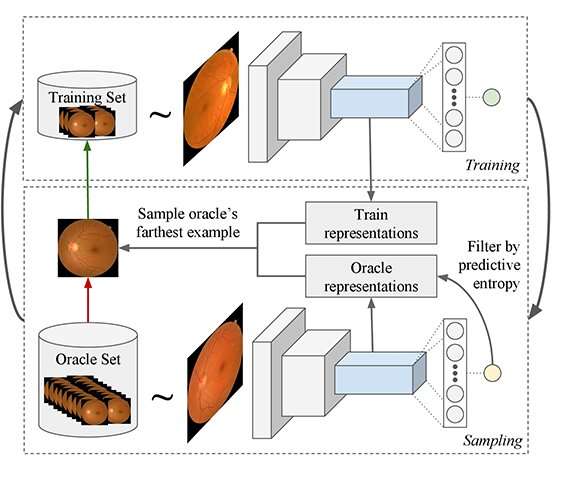
The researchers' model begins working with a set of unlabeled images. The model decides how many images to label to have a robust and accurate set of training data. It chooses an initial set of random data to label. Once that data is labelled, it plots that data over a distribution because the images will vary by age, gender, physical property, etc. In order to make a good decision based on this data, the samples need to cover a large distribution space. The system then decides what new data should be added to the dataset, considering the current distribution of data.
"The system measures how optimal this distribution is," said Noh, an associate professor of civil and environmental engineering, "and then computes metrics when a certain set of new data is added to it, and selects the new dataset that maximizes its optimality."
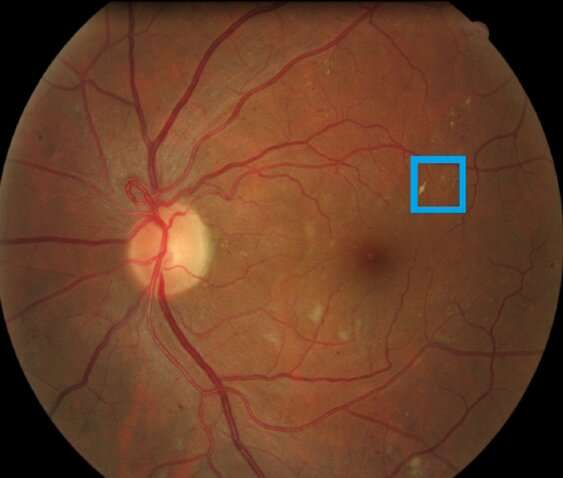
The process is repeated until the set of data has a good enough distribution to be used as the training set. Their method, called MedAL (for medical active learning), achieved 80% accuracy on detecting diabetic retinopathy, using only 425 labeled images, which is a 32% reduction in the number of required labeled examples compared to the standard uncertainty sampling technique, and a 40% reduction compared to random sampling.
They also tested the model on other diseases, including skin cancer and breast cancer images, to show that it could apply to a variety of different medical images. The method is generalizable, since its focus is on how to use data strategically rather than trying to find a specific pattern or feature for a disease. It could also be applied to other problems that use deep learning but have data constraints.
"Our active learning approach combines predictive entropy-based uncertainty sampling and a distance function on a learned feature space to optimize the selection of unlabeled samples," said Smailagic, a research professor in Carnegie Mellon's Engineering Research Accelerator. "The method overcomes the limitations of the traditional approaches by efficiently selecting only the images that provide the most information about the overall data distribution, reducing computation cost and increasing both speed and accuracy."
The team included civil and environmental engineering Ph.D. students Mostafa Mirshekari, Jonathon Fagert, and Susu Xu, and electrical and computer engineering master's students Devesh Walawalkar and Kartik Khandelwal. They presented their findings at the 2018 IEEE International Conference on Machine Learning and Applications in December, where they received a Best Paper Award for their novel work.
More information: Asim Smailagic et al. MedAL: Accurate and Robust Deep Active Learning for Medical Image Analysis, 2018 17th IEEE International Conference on Machine Learning and Applications (ICMLA) (2019). DOI: 10.1109/ICMLA.2018.00078