Recognition software drives matches across multiple science domains

The world is awash in images. Current estimates are that there were 2.1 billion smart phone users in mid-2016, up by half a billion since mid-2014, and they are generating a tremendous number of photos. In a perfect world, no one would store a photo without carefully annotating the place, time, and content next to the image. Of course, most of us are too busy grabbing new images to carefully curate the existing ones.
The same is often true for scientists at work, who collect images without always annotating all the details. Moreover, the vast majority of scientific images are taken automatically, at ever faster rates, with detectors, computers and bandwidth reaching unprecedented acquisition levels.
Annotating these images as they are collected isn't feasible—they show up far too fast to be analyzed and catalogued. And sometimes it's hard to know exactly what an image is, until it is compared to others.
Now, the same technology that helps online shoppers search for similar shoes or lamps also holds promise for helping scientists better store, analyze and compare images from experiments. Scientists at the Department of Energy's Lawrence Berkeley National Laboratory (Berkeley Lab), working jointly with colleagues from the Berkeley Institute for Data Science at UC Berkeley, are building automatic machine learning tools to make inferences and recognize characteristics in images generated by experiments ranging from cells to composite materials.
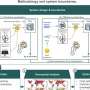
The team, led by Daniela Ushizima of Berkeley Lab's Computational Research Division, has built a new Python-based tool for content-based image retrieval (CBIR) capable of searching relevant items from large datasets, given unseen samples. Named "pyCBIR," the tool can be used to catalog and retrieve images from different science domains, such as biology, materials research and geology. The concept is analogous to the process behind tagging photos of friends in Facebook – after explicitly marking photos as belonging to the same person a number of times, Facebook "learns" to apply the tag to new photo posts of the same person.
Unlike pictures of faces which almost all have the same basic landmarks, scientific images from experiments typically present a much wider range of properties, which even experts may have difficulty deciphering. This makes the automated recognition process even more complex. To develop a method that would work with so many different characteristics, Ushizima and her team had to conduct a number of experiments themselves.
To transform the raw picture into a set of signatures, the team ran the images through layers of processing using a family of algorithms known as convolutional neural networks. The goal was to create several data-driven models that enable automated assignment of patterns, e.g., cells, to different classes using scientific images they were interested in. By repeating the process over and over, they "trained" the application to improve its accuracy.
"We looked at the signatures of shapes, including image color and texture and then how to turn that information into numbers so we could index and catalog different patterns," said Ushizima, who is also a fellow at the Berkeley Institute for Data Science at UC Berkeley and a member of Berkeley Lab's Center for Advanced Mathematics for Energy Research Applications (CAMERA). Ushizima is finding interest from other organizations, and intends to soon make pyCBIR publicly available. The core ideas inside pyCBIR were presented in a talk at the2016 PyData Conference held August 12-14 in San Francisco, but lots of additional developments are outlined in a new article submitted for publication.
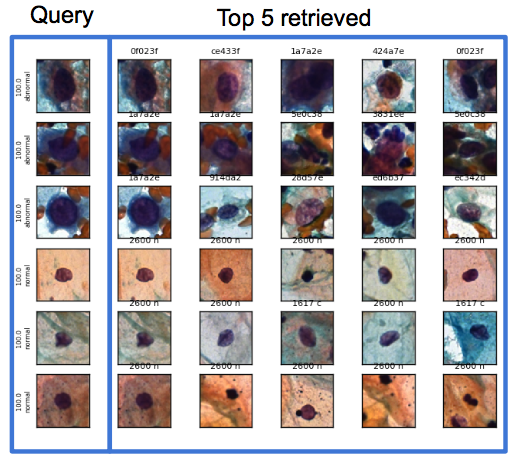
"The challenge was how to represent these signatures to reflect distances, angles and borders in the images," Ushizima said. "The idea was to come up with a way to understand what's similar to what else, particularly useful when dealing with data that's very abstract. pyCBIR lets you query large datasets using an image, and automatically determine the relevance to the previously catalogued images based on their similarity."
Ushizima used part of her 2015 DOE Early Career Research Program award to kick start the algorithmic project, with the goal of making a tool that can reach across scientific domains, aggregating value to data collected at the Department of Energy imaging facilities and beyond.

Getting Up to Speed
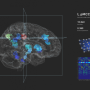
One of the first use of the underlying algorithms was in search databases of cell images, including those related to cervical cancer. The cervical cancer screening routine currently relies on two or more cytologists looking at the images under a microscope, but if this could be done at higher speed and accurately, then doctors could identify cancerous cells much earlier, which would provide more options for treatment and improve chances of survival.
A typical Pap smear contains anywhere from 50,000 to 300,000 cells, and in some cases a single abnormal cell might be lurking amongst them. Different cellular shapes, overlapping cells, poor contrast of the cytoplasm, and the presence of blood, inflammatory cells and mucus, can further complicate the analysis and lead to false negatives.
At the IEEE International Symposium on Biomedical Imaging (ISBI 2014) in Beijing, China, a method developed by Ushizima in collaboration with Professors Andrea Bianchi and Claudia Carneiro of the Federal University of Ouro Preto (UFOP) in Brazil was judged to be the fastest and most accurate in a competition between software tools to extract the boundaries of individual cytoplasm and nucleus from overlapping cervical cell images.
Working Directly with Scientists
The team has brought together a range of scientific domains, including Advanced Light Source materials scientists Alexander Hexemer and Dula Parkinson, Science without Borders visiting computer science Professors Flavio Araujo and Romuere Silva from the Federal University of Ceara in Brazil, and cervical cell researchers from the University of Manchester in England and the Federal University of Ouro Preto in Brazil.
Silva said the team is already working in the next version of pyCBIR with the goal of reducing the time to retrieve images and improving the graphical interface. "In this way, we hope to make this tool even more useful for scientists," he said.
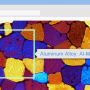
One of her collaborations has been with materials scientists working at the Advanced Light Source, a DOE Office of Science national user facility at Berkeley Lab. One ALS experiment involves putting composite ceramic materials under increasing pressure until cracks begin to form. Part of the experiment is to embed fibers in the materials to help resist cracking. The work is done in collaboration with Prof. Robert Ritchie, a professor at UC Berkeley who has been studying material failures for more than 30 years.
"Typically scientists would rely on post-docs to manually go through the images, looking for the location of fibers and identifying them one by one" says Parkinson. "We hope to use pyCBIR to leverage the invested manual labor and create models based on human-curated data. And, as the next step, pyCBIR could also streamline the effort needed to conduct the experiments by suggesting fiber locations automatically. We often put pressure on the material until it breaks, and that requires round-the-clock observation. But if we knew when and where it would break, we could focus the beamline precisely on that area and get higher-resolution images, which would give us more information."
Community Building Aspect
The future might bring a 'Facebook' of science images and help connect researchers by identifying materials and features they 'know' in common. For example, scientists sharing a particular microscope, or working across multiple beamlines, might be able to merge information about the structures in images they are all collecting, without sharing their data per se, and common characteristics might be detected and identified.
"Our prototype tool solves some problems in image search, and we will move forward by helping to recognize patterns for scientists who rely on studying specific pictures as part of their investigations," Ushizima said. "Collaboration has been vital to developing these tools which we believe will benefit a larger community."