Machine learning for damaging mutations prediction

Next-generation sequencing technology has ushered in a new era in medicine, making it easier to identify a sequence of nucleotides in DNA or a sequence of amino acids in the proteins of a specific individual and use this information for diagnosis and treatment. Minute alterations in these sequences can be indicative of a minor disorder, and sometimes a grave disease.
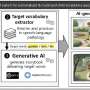
Scientists from Skoltech, the Technical University of Munich, St. Petersburg Polytechnic University and the Indian Institute of Technology Madras (Chennai, India) have developed a machine-learning-based method to analyze the atomic structures of proteins and predict the pathogenicity of mutations. The method is adapted for transmembrane proteins that account for 25 to 30 percent of all the proteins in a cell, and often serve as targets for drugs.
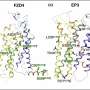
"In this study, we used a combination of 1-D information on the amino acid sequences of proteins and 3-D information on the protein's atomic structures to create an effective machine-learning-based model that helps identify disease-associated amino acid substitutions in membrane proteins," says the first author of the study and assistant professor at Skoltech, Petr Popov.
More information: Petr Popov et al, Prediction of disease-associated mutations in the transmembrane regions of proteins with known 3D structure, PLOS ONE (2019). DOI: 10.1371/journal.pone.0219452