Visual analytics tool plucks elusive patterns from elaborate datasets
From materials science and earth system modeling to quantum information science and cybersecurity, experts in many fields run simulations and conduct experiments to collect the abundance of data necessary for scientific progress. But gleaning useful insights from those data can be a challenge, especially when multiple complex variables influence research results.
To better analyze the so-called multivariate data, researchers at the Department of Energy's Oak Ridge National Laboratory developed an open-source, customizable visual analytics system called CrossVis. Unlike similar tools—which tend to focus on numerical data and provide a single visual representation of results—CrossVis juggles numerical, categorical and image-based data while providing multiple dynamic, coordinated views of these and other data types.
ORNL researchers John Goodall, Junghoon Chae, Artem Trofimov and Chad Steed, director of the ORNL Visual Informatics for Science and Technology Advances, or VISTA, laboratory, made CrossVis available online and published the system's unique capabilities in Graphics and Visual Computing.
"CrossVis is a one-stop shop for analyzing many different types of data, and it reveals relationships among more than just two variables," Steed said.
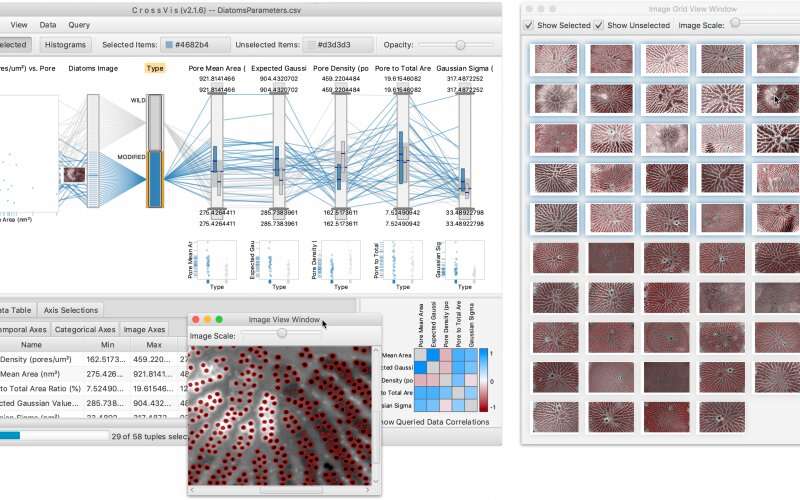
The tool's main view consists of a parallel coordinates plot, or PCP, which is a popular information visualization technique. PCPs display a data table's columns as vertical axes and its rows as polylines, which are chains of interdependent line segments connected to the axes. In this case, the CrossVis interface extends beyond traditional PCPs to include nonnumerical data, which have no natural order, and temporal, or time-based, data.
Additionally, CrossVis provides scatterplots, image panes and other options that complement the main view to help users identify key patterns and interesting anomalies in heterogenous, multivariate data. To narrow their focus, users can also choose to highlight a variable in all views simultaneously, generate new data or input parameters to filter existing data.
"Before, scientists had to use individual programs to analyze image data, numerical data and categorical data, then manually compare the results," Steed said. "CrossVis lets them complete all those steps within a single framework."
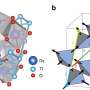
The team took advantage of the system's ability to analyze categorical and image data by applying it to a genetic engineering project led by researchers at ORNL's Center for Nanophase Materials Sciences, or CNMS, which involved verifying results from an artificial neural network, or ANN, applied to scanning electron microscopy images of diatoms. A type of algae, diatoms produce strong silica that could be useful for industrial purposes, including drug delivery and water filtration.
Specifically, the CNMS team characterized pores on the diatoms to distinguish between unmodified, or wild, diatoms and genetically modified versions of these organisms. Eventually, these insights could help scientists optimize and emulate diatom biomineralization, which is the process these organisms use to generate silica.
The team used CrossVis to examine relationships between diatom parameters, and the tool's many views revealed subtle differences between the two categories. For example, the researchers determined that wild diatoms have more pores that are smaller than those of their modified counterparts, which have fewer pores that are larger in size.
"The ANN automatically derived image classifications that identified pores as an important feature for separating the two types of diatoms," Steed said. "However, these results didn't clearly show why the algorithm chose to classify pores the way it did, so CrossVis enabled the CNMS scientists to interpret and verify their findings."
"Without CrossVis, we would not as thoroughly understand how to differentiate between wild and modified diatom images based on these crucial parameters, namely mean area and the density of pores," added ORNL researcher Artem Trofimov, who led the CNMS project.
To prove the value of CrossVis at a larger scale, Steed and his collaborators also worked with the ORNL-led team that developed the Energy Exascale Earth System Model to help validate climate modeling techniques. Additionally, the team used CrossVis to verify data in the National Oceanic and Atmospheric Administration's Atlantic Hurricane Database, which contains 21 columns and more than 50,000 rows of statistical information about the locations, sizes and other characteristics of hurricanes over time.
"That was a good use case because it was a much larger dataset with more variables," Steed said. "We found patterns that confirmed known hurricane conditions, which demonstrated that CrossVis can effectively validate real-world results on a larger scale."
Going forward, the CrossVis team aims to further improve this resource. For example, the researchers plan to scale up CrossVis to run on high-performance computing systems. With the processing power of supercomputers, such as ORNL's Summit, CrossVis could more efficiently complete complex calculations.
By incorporating automated machine learning techniques, the team plans to more actively capture user interactions with the data. Scientists would label data samples, and built-in artificial intelligence algorithms would then identify, label and compile similar patterns in unseen sections of the data, enabling users to quickly analyze entire datasets and potentially make unexpected discoveries.
"If you tried to sort through something like the hurricane dataset or climate modeling data manually, it would take a lifetime," Steed said. "This kind of human-machine cooperation, which combines the creativity and intuition of domain experts with the data-crunching power of computers, is the key to more effective data analysis."
More information: Chad A. Steed et al. CrossVis: A visual analytics system for exploring heterogeneous multivariate data with applications to materials and climate sciences, Graphics and Visual Computing (2020). DOI: 10.1016/j.gvc.2020.200013