March 30, 2021 feature
MolMapNet: An out-of-the-box deep learning model to predict pharmaceutical properties

Over the past few decades, computer scientists have developed deep learning tools for a broad variety of applications, including for the analysis of pharmaceutical drugs. Most recently, deep learning models that predict the properties of pharmaceuticals have been trained to analyze and learn molecular representations.
Researchers at Tsinghua University, the National University of Singapore, Fudan University's School of Pharmacy, and Zheijang University have recently developed MolMapNet, a new artificial intelligence (AI) tool that can predict the pharmaceutical properties of drugs by analyzing human-knowledge-based molecular representations. This tool, presented in a paper published in Nature Machine Intelligence, can also be used by people with little or no knowledge of computer science, biology or other sciences.
"We were aware that pharmaceutical investigations require the learning of many molecular characters, particularly the rich collection of molecular properties (like volume) derived from human knowledge, but these molecular properties are tough to learn by AI (artificial intelligence)," Yu Zong Chen, one of the researchers who carried out the study, told TechXplore.
While AI tools are generally good at recognizing images that are spatially ordered (e.g., images of objects), they do not perform as well on unordered data such as molecular properties. This characteristic significantly impairs their performance on the analysis of pharmaceuticals. Chen and his colleagues wanted to overcome this limitation in order to improve the performance of deep-learning models for predicting pharmaceutical properties.
"With limited pharmaceutical data, it is hard to improve AI architectures," Chen said. "We asked whether we could improve the way AI reads molecular properties. Our solution is to map unordered molecular properties into ordered images for AI to more efficiently recognize molecular properties."
This innovative out-of-the-box AI tool does not require parameter fine tuning, which means that it is also accessible to non-expert users. Remarkably, the researchers found that it outperformed state-of-the-art AI tools on most of the 26 pharmaceutical benchmark datasets.
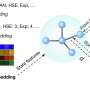
"Our approach follows three steps for improved deep learning prediction of pharmaceutical properties," Chen said. "The first step is to broadly learn the intrinsic relationships of molecular properties from over 8 million molecules. These relationships may be linked to and thus indicators of various pharmaceutical properties."
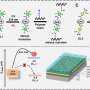
The second step of the approach entails the use of a newly developed data transformation technique to map the molecular properties of pharmaceuticals into 2D images, where the pixel layouts reflect the intrinsic relationships between these properties. These pixel layouts contain crucial indicators of pharmaceutical properties that can be captured by adequately trained deep learning models.
As a third step, the researchers trained an image-recognition tool to learn the 2D images and use them to predict pharmaceutical properties. The AI tool can capture specific pixel layout patterns that characterize specific pharmaceutical properties, similarly to how AI techniques might discern between males and females in a picture by looking at hair length or other gender-related features.
"There are two notable achievements of our study," Chen said. "The first is the introduction of a new method for mapping unordered molecular properties into ordered images that present the intrinsic relationships of molecular properties. The second is the development of an innovative out-of-the-box AI tool for deep-learning prediction of pharmaceutical properties by non-experts with state-of-the-art performance."
In the future, the out-of-the-box deep learning model could significantly speed up pharmaceutical research, helping scientists to predict the properties of different drugs faster and more efficiently. In their next studies, Chen and his colleagues plan to develop their model further, so that it can also be applied to biomedical studies.
More information: Out-of-the-box deep learning prediction of pharmaceutical properties by broadly learned knowledge-based molecular representations. Nature Machine Intelligence(2021). DOI: 10.1038/s42256-021-00301-6
© 2021 Science X Network