Deep-learning–based image analysis is now just a click away
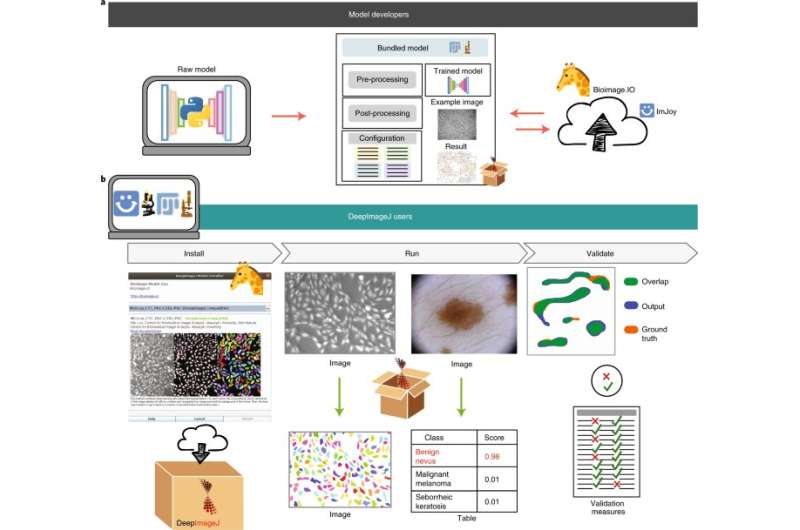
Under an initiative by EPFL's Center for Imaging, a team of engineers from EPFL and Universidad Carlos III de Madrid have developed a plugin that makes it easier to incorporate artificial intelligence into image analysis for life-science research. The plugin, called deepImageJ, is described in a paper appearing today in Nature Methods.
Over the past five years, image analysis has been shifting away from traditional mathematical- and observational-based methods towards data-driven processing and artificial intelligence. This major development is making the detection and identification of valuable information in images easier, faster, and increasingly automated—in just about every research field. When it comes to life science, deep-learning-, a sub-field of artificial intelligence, is showing an increasing potential for bioimage analysis. Unfortunately, using the deep-learning models often requires coding skills that few life scientists possess. To make the process easier, image analysis experts from EPFL and UC3M, working in association with EPFL's Center for Imaging, have developed deepImageJ—an open-source plugin that's described in a paper published today in Nature Methods.
Using neural networks in biomedical research
Deep-learning models are a significant breakthrough for the many fields that rely on imaging, such as diagnostics and drug development. In bio-imaging, for example, deep learning can be used to process vast collections of images and detect lesions in organic tissue, identify synapses between nerve cells, and determine the structure of cell membranes and nuclei. It's ideal for recognizing and classifying images, identifying specific elements, and predicting experimental results.
This type of artificial intelligence involves training a computer to perform a task by drawing on large amounts of previously annotated data. It's similar to CCTV systems that perform facial recognition, or to mobile-camera apps that enhance photos. Deep-learning models are based on sophisticated computational architectures called artificial neural networks that can be trained for specific research purposes, such as to recognize certain types of cells or tissue lesions or to improve image quality. The trained neural network is then saved as a computer model.
Artificial intelligence, but without the code
For biomedical imaging, a consortium of European researchers is developing a repository of these pre-trained deep-learning models, called the BioImage Model Zoo. "To train these models, researchers need specific resources and technical knowledge—especially in Python coding—that many life scientists do not have," says Daniel Sage, the engineer at EPFL's Center for Imaging who is overseeing the deepImageJ development. "But ideally, these models should be available to everyone."
The deepImageJ plugin bridges the gap between artificial neural networks and the researchers who use them. Now, a life scientist can ask a computer engineer to design and train a machine-learning algorithm to perform a specific task, which the scientist can then easily run via a user interface—without ever seeing a single line of code. The plugin is open-source and free-of-charge, and will speed the dissemination of new developments in computer science and the publication of biomedical research. It is designed to be a collaborative resource that enables engineers, computer scientists, mathematicians and biologists to work together more efficiently. For example, a model developed recently by an EPFL Master's student, working as part of a cross-disciplinary team, enables scientists to distinguish human cells from mouse cells in tissue sections.
Researchers can train users, too
Life scientists around the world have been hoping for such a system for several years, but—until EPFL's Center for Imaging stepped in—no one had taken up the challenge of building one. The research group is headed by Daniel Sage and Michael Unser, the Center's academic director, together with Arrate Muñoz Barrutia, associate professor at UC3M. Professor Muñoz-Barrutia led the operational development work along with one of her Ph.D. students, Estibaliz Gómez-de-Mariscal, and Carlos García López de Haro, a bioengineering research assistant .
So that as many researchers can use the plugin as possible, the group is also developing virtual seminars, training materials and online resources, with a view to better exploiting the full potential of artificial intelligence. These materials are being designed with both programmers and life scientists in mind, so that users can quickly come to grips with the new method. DeepImageJ will also be presented at ZIDAS—a week-long class on image and data analysis for life scientists in Switzerland.
More information: Estibaliz Gómez-de-Mariscal et al, DeepImageJ: A user-friendly environment to run deep learning models in ImageJ, Nature Methods (2021). DOI: 10.1038/s41592-021-01262-9