Deep learning accelerates the detection of live bacteria using thin-film transistor arrays
Early detection and identification of pathogenic bacteria in food and water samples are essential to public health. Bacterial infections cause millions of deaths worldwide and bring a heavy economic burden, costing more than 4 billion dollars annually in the United States alone. Among pathogenic bacteria, Escherichia coli (E. coli) and other coliform bacteria are among the most common ones, and they indicate fecal contamination in food and water samples. The most conventional and frequently used method for detecting these bacteria involves culturing of the samples, which usually takes >24 hours for the final read-out and needs expert visual examination. Although some methods based on, for example, the amplification of nucleic acids, can reduce the detection time to a few hours, they cannot differentiate live and dead bacteria and present low sensitivity at low concentrations of bacteria. That is why the U.S. Environmental Protection Agency (EPA) approves no nucleic acid-based bacteria sensing method for screening water samples.
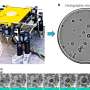
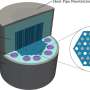
In an article recently published in ACS Photonics, a journal of the American Chemical Society (ACS), a team of scientists, led by Professor Aydogan Ozcan from the Electrical and Computer Engineering Department at the University of California, Los Angeles (UCLA), and co-workers have developed an AI-powered smart bacterial colony detection system using a thin-film transistor (TFT) array, which is a widely used technology in mobile phones and other displays.
The ultra-large imaging area of the TFT array (27 mm × 26 mm) manufactured by researchers at Japan Display Inc. enabled the system to rapidly capture the growth patterns of bacterial colonies without the need for scanning, which significantly simplified both the hardware and software design. This system achieved ~12-hour time savings compared to gold-standard culture-based methods approved by EPA. By analyzing the microscopic images captured by the TFT array as a function of time, the AI-based system could rapidly and automatically detect colony growth with a deep neural network. Following the detection of each colony, a second neural network is used to classify the bacteria species.
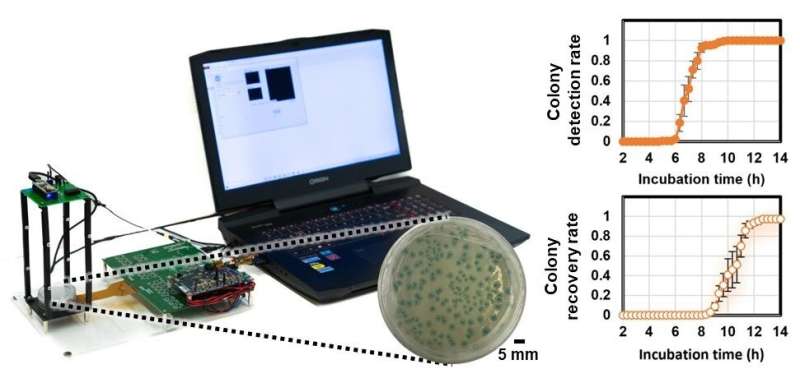
The efficacy of this automated bacterial colony detection system was demonstrated by performing early detection and classification of three types of bacteria, i.e., E. coli, Citrobacter, and Klebsiella pneumoniae (K. pneumoniae). The researchers achieved a colony detection rate of >90% within nine hours and further identified their species at ∼12 hours, corresponding to a time saving of ~12 hours compared to the EPA-approved culture methods. In addition, all the digital processing steps take <25 sec using a standard computer without needing an advanced graphics processing unit (GPU).
These results demonstrate the feasibility of this automated, AI-based bacterial colony detection system using TFT arrays as a rapid, cost-effective, and accurate technique, which is especially suitable for resource-limited environments. Due to the low-cost, low-heat generation, large scalability, and low power consumption of TFT arrays widely used in mobile displays, this automated colony detection platform has massive potential to be used in microbiology research and field-based bacteria sensing.
More information: Yuzhu Li et al, Deep Learning-Enabled Detection and Classification of Bacterial Colonies Using a Thin-Film Transistor (TFT) Image Sensor, ACS Photonics (2022). DOI: 10.1021/acsphotonics.2c00572