This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
proofread
New neural network method improves microscopic distance measurements between colored points in three dimensions
Fluorescence microscopy is a widely used technique in the life sciences that enables scientists to see specific parts of cells and tissues by labeling them with glowing molecules, helping in the study of cell structure and movement, molecule behavior, and drug effects.
To understand the relative position of structures within a cell or tissue, scientists normally must compute the 3-dimensional (3D) distance between two microscopic fluorescent spots using position information. However, scientists from Technion–Israel Institute of Technology leveraged neural networks to design an optical system to measure the distance directly.
The research was published in Intelligent Computing on Dec. 21.
"Microscopes have been traditionally optimized to output the sharpest possible image, targeted for the human observer. However, in computational microscopy, this goal has changed toward providing the most informative measurement for subsequent computer analysis," the authors said. The use of computational techniques can enhance the capabilities of traditional optical microscopes.
One of the important goals of contemporary microscopy is confirming the 3D spatial position of fluorescent sources. The resolution of optical microscopy, and thus the accuracy of position measurements, is limited by the diffraction, or the "spreading out," of the light wave when it passes through a small aperture or is focused to a tiny spot. This spot, referred to as the point spread function (PSF) of the system, can be engineered to encode useful characteristics of a point source, such as color information, or 3D spatial position for accurate localization.
However, in many multicolor localization imaging scenarios, scientists need to measure the 3D distance between two spots in the sample, rather than determine their precise locations. The authors illustrate this point: "When the sample consists of precisely 2 differently colored spots, e.g., labeled DNA loci in live yeast cells, the precise 3D position of each locus with respect to some global coordinate system is irrelevant—only the distance between the 2 loci is of interest."
Accurate 3D distance measurements can reveal the movement and dynamics of components over time, aiding in the study of cellular processes such as migration and division.
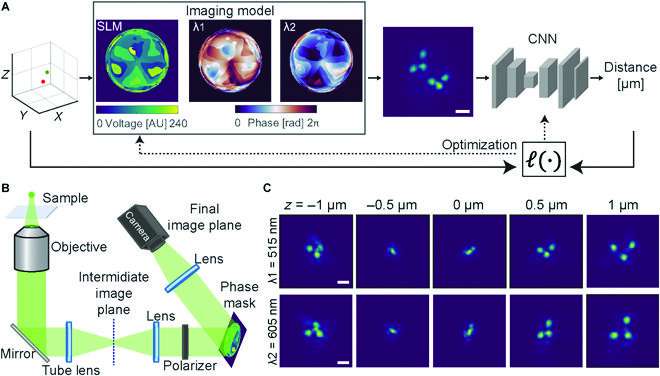
In order to solve the problem of obtaining such measurements, the authors developed an end-to-end machine-learning method of optical system design that combines neural networks and PSF engineering to directly measure the 3D distance between two differently-colored spots. A neural network, comprised of a physical model and a reconstruction network, is being used to learn simultaneously both the encoding (the optimal phase mask for the PSF modulation) and the decoding (the computation of the distance from the modified images).
The authors demonstrated experimentally, not only by using pairs of fluorescent beads, but also by using pairs of DNA loci in yeast cells, that it is possible to use neural networks in combination with PSF engineering to design a more accurate optical system that takes advantage of the need for distance measurements rather than position measurements.
Future work could include single-color distance determination, for labeling simplification and improved signal-to-noise ratio (SNR), or rather expanding this method to the encoding of more than two emission colors and investigating more complex dynamics, specifically for chromosome organization research. In addition, the method created by these researchers could be further improved to be used to process images containing crowded cells for high-throughput imaging of cell populations.
More information: Ofri Goldenberg et al, Learning Optimal Multicolor PSF Design for 3D Pairwise Distance Estimation, Intelligent Computing (2022). DOI: 10.34133/icomputing.0004