This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Deep neural network provides robust detection of disease biomarkers in real time

Sophisticated systems for the detection of biomarkers—molecules such as DNA or proteins that indicate the presence of a disease—are crucial for real-time diagnostic and disease-monitoring devices.
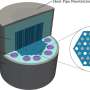
Holger Schmidt, distinguished professor of electrical and computer engineering at UC Santa Cruz, and his group have long been focused on developing unique, highly sensitive devices called optofluidic chips to detect biomarkers.
Schmidt's graduate student Vahid Ganjalizadeh led an effort to use machine learning to enhance their systems by improving its ability to accurately classify biomarkers. The deep neural network he developed classifies particle signals with 99.8 percent accuracy in real time, on a system that is relatively cheap and portable for point-of-care applications, as shown in a new paper in Scientific Reports.
When taking biomarker detectors into the field or a point-of-care setting such as a health clinic, the signals received by the sensors may not be as high quality as those in a lab or a controlled environment. This may be due to a variety of factors, such as the need to use cheaper chips to bring down costs, or environmental characteristics such as temperature and humidity.
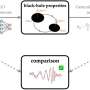
To address the challenges of a weak signal, Schmidt and his team developed a deep neural network that can identify the source of that weak signal with high confidence. The researchers trained the neural network with known training signals, teaching it to recognize potential variations it could see, so that it can recognize patterns and identify new signals with very high accuracy.
First, a parallel cluster wavelet analysis (PCWA) approach designed in Schmidt's lab detects that a signal is present. Then, the neural network processes the potentially weak or noisy signal, identifying its source. This system works in real time, so users are able to receive results in a fraction of a second.
"It's all about making the most of possibly low quality signals, and doing that really fast and efficiently," Schmidt said.
A smaller version of the neural network model can run on portable devices. In the paper, the researchers run the system over a Google Coral Dev board, a relatively cheap edge device for accelerated execution of artificial intelligence algorithms. This means the system also requires less power to execute the processing compared to other techniques.
"Unlike some research that requires running on supercomputers to do high-accuracy detection, we proved that even a compact, portable, relatively cheap device can do the job for us," Ganjalizadeh said. "It makes it available, feasible, and portable for point-of-care applications."
The entire system is designed to be used completely locally, meaning the data processing can happen without internet access, unlike other systems that rely on cloud computing. This also provides a data security advantage, because results can be produced without the need to share data with a cloud server provider.
It is also designed to be able to give results on a mobile device, eliminating the need to bring a laptop into the field.
"You can build a more robust system that you could take out to under-resourced or less- developed regions, and it still works," Schmidt said.
This improved system will work for any other biomarkers Schmidt's lab's systems have been used to detect in the past, such as COVID-19, Ebola, flu, and cancer biomarkers. Although they are currently focused on medical applications, the system could potentially be adapted for the detection of any type of signal.
To push the technology further, Schmidt and his lab members plan to add even more dynamic signal processing capabilities to their devices. This will simplify the system and combine the processing techniques needed to detect signals at both low and high concentrations of molecules. The team is also working to bring discrete parts of the setup into the integrated design of the optofluidic chip.
More information: Vahid Ganjalizadeh et al, Machine learning at the edge for AI-enabled multiplexed pathogen detection, Scientific Reports (2023). DOI: 10.1038/s41598-023-31694-6