From one brain scan, more information for medical artificial intelligence
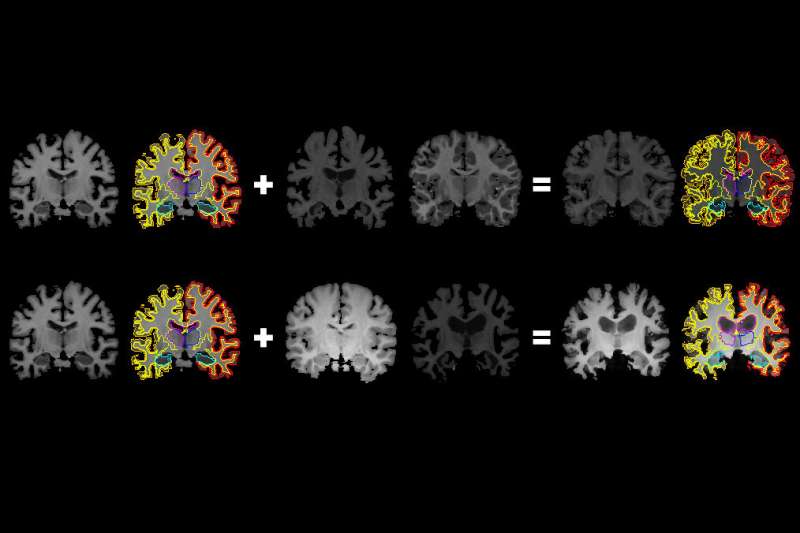
MIT researchers have devised a novel method to glean more information from images used to train machine-learning models, including those that can analyze medical scans to help diagnose and treat brain conditions.
An active new area in medicine involves training deep-learning models to detect structural patterns in brain scans associated with neurological diseases and disorders, such as Alzheimer's disease and multiple sclerosis. But collecting the training data is laborious: All anatomical structures in each scan must be separately outlined or hand-labeled by neurological experts. And, in some cases, such as for rare brain conditions in children, only a few scans may be available in the first place.
In a paper presented at the recent Conference on Computer Vision and Pattern Recognition, the MIT researchers describe a system that uses a single labeled scan, along with unlabeled scans, to automatically synthesize a massive dataset of distinct training examples. The dataset can be used to better train machine-learning models to find anatomical structures in new scans—the more training data, the better those predictions.
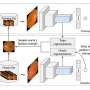
The crux of the work is automatically generating data for the "image segmentation" process, which partitions an image into regions of pixels that are more meaningful and easier to analyze. To do so, the system uses a convolutional neural network (CNN), a machine-learning model that's become a powerhouse for image-processing tasks. The network analyzes a lot of unlabeled scans from different patients and different equipment to "learn" anatomical, brightness, and contrast variations. Then, it applies a random combination of those learned variations to a single labeled scan to synthesize new scans that are both realistic and accurately labeled. These newly synthesized scans are then fed into a different CNN that learns how to segment new images.
"We're hoping this will make image segmentation more accessible in realistic situations where you don't have a lot of training data," says first author Amy Zhao, a graduate student in the Department of Electrical Engineering and Computer Science (EECS) and Computer Science and Artificial Intelligence Laboratory (CSAIL). "In our approach, you can learn to mimic the variations in unlabeled scans to intelligently synthesize a large dataset to train your network."
There's interest in using the system, for instance, to help train predictive-analytics models at Massachusetts General Hospital, Zhao says, where only one or two labeled scans may exist of particularly uncommon brain conditions among child patients.
Joining Zhao on the paper are: Guha Balakrishnan, a postdoc in EECS and CSAIL; EECS professors Fredo Durand and John Guttag, and senior author Adrian Dalca, who is also a faculty member in radiology at Harvard Medical School.
The "Magic" behind the system
Although now applied to medical imaging, the system actually started as a means to synthesize training data for a smartphone app that could identify and retrieve information about cards from the popular collectable card game, "Magic: The Gathering." Released in the early 1990s, "Magic" has more than 20,000 unique cards—with more released every few months—that players can use to build custom playing decks.
Zhao, an avid "Magic" player, wanted to develop a CNN-powered app that took a photo of any card with a smartphone camera and automatically pulled information such as price and rating from online card databases. "When I was picking out cards from a game store, I got tired of entering all their names into my phone and looking up ratings and combos," Zhao says. "Wouldn't it be awesome if I could scan them with my phone and pull up that information?"
But she realized that's a very tough computer-vision training task. "You'd need many photos of all 20,000 cards, under all different lighting conditions and angles. No one is going to collect that dataset," Zhao says.
Instead, Zhao trained a CNN on smaller dataset of around 200 cards, with 10 distinct photos of each card, to learn how to warp a card into various positions. It computed different lighting, angles, and reflections—for when cards are placed in plastic sleeves—to synthesized realistic warped versions of any card in the dataset. It was an exciting passion project, Zhao says: "But we realized this approach was really well-suited for medical images, because this type of warping fits really well with MRIs."
Mind warp
Magnetic resonance images (MRIs) are composed of three-dimensional pixels, called voxels. When segmenting MRIs, experts separate and label voxel regions based on the anatomical structure containing them. The diversity of scans, caused by variations in individual brains and equipment used, poses a challenge to using machine learning to automate this process.
Some existing methods can synthesize training examples from labeled scans using "data augmentation," which warps labeled voxels into different positions. But these methods require experts to hand-write various augmentation guidelines, and some synthesized scans look nothing like a realistic human brain, which may be detrimental to the learning process.
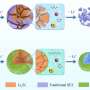
Instead, the researchers' system automatically learns how to synthesize realistic scans. The researchers trained their system on 100 unlabeled scans from real patients to compute spatial transformations—anatomical correspondences from scan to scan. This generated as many "flow fields," which model how voxels move from one scan to another. Simultaneously, it computes intensity transformations, which capture appearance variations caused by image contrast, noise, and other factors.
In generating a new scan, the system applies a random flow field to the original labeled scan, which shifts around voxels until it structurally matches a real, unlabeled scan. Then, it overlays a random intensity transformation. Finally, the system maps the labels to the new structures, by following how the voxels moved in the flow field. In the end, the synthesized scans closely resemble the real, unlabeled scans—but with accurate labels.
To test their automated segmentation accuracy, the researchers used Dice scores, which measure how well one 3-D shape fits over another, on a scale of 0 to 1. They compared their system to traditional segmentation methods—manual and automated—on 30 different brain structures across 100 held-out test scans. Large structures were comparably accurate among all the methods. But the researchers' system outperformed all other approaches on smaller structures, such as the hippocampus, which occupies only about 0.6 percent of a brain, by volume.
"That shows that our method improves over other methods, especially as you get into the smaller structures, which can be very important in understanding disease," Zhao says. "And we did that while only needing a single hand-labeled scan."
More information: Data augmentation using learned transformations for one-shot medical image segmentation arXiv:1902.09383 [cs.CV] arxiv.org/abs/1902.09383
This story is republished courtesy of MIT News (web.mit.edu/newsoffice/), a popular site that covers news about MIT research, innovation and teaching.