May 10, 2021 feature
SurvNet: A backward elimination procedure to enhance variable selection for deep neural networks
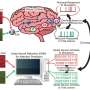
In recent years, models based on deep neural networks have achieved remarkable results on numerous tasks. Despite their high prediction accuracy, these models are known for their "black-box" nature, which essentially means that the processes that lead to their predictions are difficult to interpret.
One of the key processes that a deep neural network performs when learning to make predictions is known as variable selection. Essentially, this entails the selection of input variables that have a strong predictive power (i.e., the identification of data features that allow a model to make highly accurate predictions).
Researchers at University of Notre Dame recently developed SurvNet, a technique that could improve variable selection processes when training deep neural networks. This technique, presented in a paper published in Nature Machine Intelligence, can estimate and control false discovery rates during variable selection (i.e., the extent to which a deep neural network selects variables that are irrelevant to the task it is meant to complete).
"People typically think of deep neural networks as black boxes (i.e., while they achieve high prediction accuracy, it's hard to explain why they work), and this limits their applications in fields that require interpretable models, such as biology and medicine," Jun Li, the principal investigator who conceived the study, told TechXplore. "We wanted to devise a method to interpret neural networks, particularly to know which input variables are important to the success of a network."
To improve variable selection, Li and his student Zixuan Song developed SurvNet, a backward elimination procedure that can be used to select input variables for deep neural networks reliably. Essentially, SurvNet gradually eliminates variables (i.e., data features) that are irrelevant in a particular task, ultimately identifying the ones with the highest predictive power.
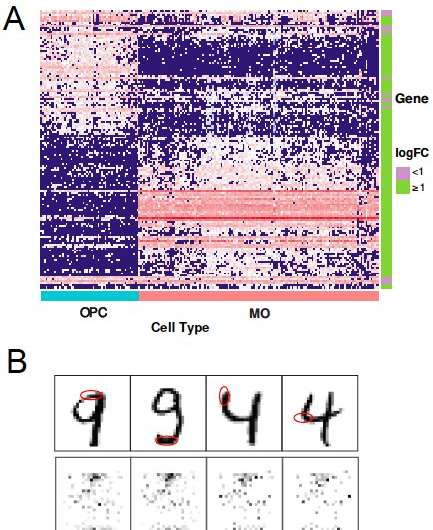
"For example, in genomics study, researchers use gene expression data, which consists of expression of thousands of genes (each gene is an input variable), to diagnose diseases," Li said. "A deep neural network may be developed for such diagnosis, but we wanted to know that which genes (typically several or dozens) are truly important for the diagnosis, so that researchers can do further experiments to study or validate these genes and learn more about the mechanisms of the disease, to finally identify chemicals/medication that tackle these genes and can cure a specific disease."
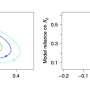
Li and Song evaluated SurvNet in a series of experiments on both real and simulated datasets. In addition, they compared its performance with that of other existing techniques for variable selection. In these tests, SurvNet compared favorably with other methods, and while some techniques (e.g., knockoff-based methods) achieved a lower false discovery rate on data with highly correlated variables, SurvNet usually had a higher variable selection power overall, achieving a better trade-off between false discoveries and power.
"The unique feature of SurvNet, is that it provides a 'quality control' for variable selection, and this quality control is done using a modern and statistically rigid way, by controlling the false discovery rate," Li said. "Such a strict quality control is pivotal for studies in biology and medicine, as further (experimental) validations of the results are often costly and time consuming."
Compared to other variable selection methods, SurvNet is more reliable and computationally efficient. In the future, it could help to improve the prediction accuracy and interpretability of models based on deep neural networks, by efficiently selecting variables with a strong predictive power.
"Our study provides a handy tool to tell which input variables are important, and this tool is automatic (no human intervention is needed), reliable (enabling strict quality control), computationally efficient (low cost in computational time or resources), and versatile (applicable to a wide-variety of problems)," Li said. "In our next studies, we plan to extend SurvNet to unsupervised studies, such as clustering."
More information: Variable selection with false discovery rate control in deep neural networks. Nature Machine Intelligence(2021). DOI: 10.1038/s42256-021-00308-z.
© 2021 Science X Network