'Minimalist machine learning' algorithms analyze images from very little data
Mathematicians at the Department of Energy's Lawrence Berkeley National Laboratory (Berkeley Lab) have developed a new approach to machine learning aimed at experimental imaging data. Rather than relying on the tens or hundreds of thousands of images used by typical machine learning methods, this new approach "learns" much more quickly and requires far fewer images.
Daniël Pelt and James Sethian of Berkeley Lab's Center for Advanced Mathematics for Energy Research Applications (CAMERA) turned the usual machine learning perspective on its head by developing what they call a "Mixed-Scale Dense Convolution Neural Network (MS-D)" that requires far fewer parameters than traditional methods, converges quickly, and has the ability to "learn" from a remarkably small training set. Their approach is already being used to extract biological structure from cell images, and is poised to provide a major new computational tool to analyze data across a wide range of research areas.
As experimental facilities generate higher resolution images at higher speeds, scientists can struggle to manage and analyze the resulting data, which is often done painstakingly by hand. In 2014, Sethian established CAMERA at Berkeley Lab as an integrated, cross-disciplinary center to develop and deliver fundamental new mathematics required to capitalize on experimental investigations at DOE Office of Science user facilities. CAMERA is part of the lab's Computational Research Division.
"In many scientific applications, tremendous manual labor is required to annotate and tag images—it can take weeks to produce a handful of carefully delineated images," said Sethian, who is also a mathematics professor at the University of California, Berkeley. "Our goal was to develop a technique that learns from a very small data set."
Details of the algorithm were published Dec. 26, 2017 in a paper in the Proceedings of the National Academy of Sciences.
"The breakthrough resulted from realizing that the usual downscaling and upscaling that capture features at various image scales could be replaced by mathematical convolutions handling multiple scales within a single layer," said Pelt, who is also a member of the Computational Imaging Group at the Centrum Wiskunde & Informatica, the national research institute for mathematics and computer science in the Netherlands.
To make the algorithm accessible to a wide set of researchers, a Berkeley team led by Olivia Jain and Simon Mo built a web portal "Segmenting Labeled Image Data Engine (SlideCAM)" as part of the CAMERA suite of tools for DOE experimental facilities.

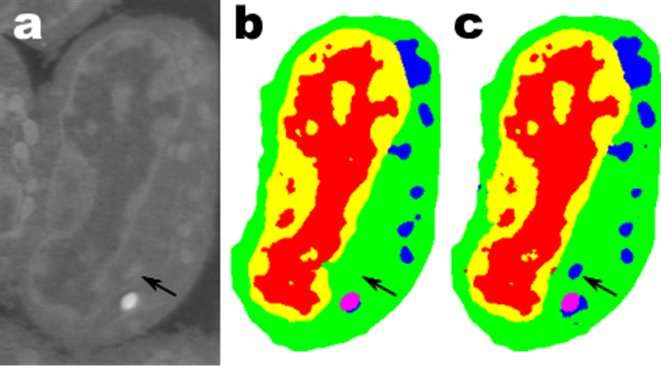
One promising application is in understanding the internal structure of biological cells and a project in which Pelt's and Sethian's MS-D method needed only data from seven cells to determine the cell structure.
"In our laboratory, we are working to understand how cell structure and morphology influences or controls cell behavior. We spend countless hours hand-segmenting cells in order to extract structure, and identify, for example, differences between healthy vs. diseased cells," said Carolyn Larabell, Director of the National Center for X-ray Tomography and Professor at the University of California San Francisco School of Medicine. "This new approach has the potential to radically transform our ability to understand disease, and is a key tool in our new Chan-Zuckerberg-sponsored project to establish a Human Cell Atlas, a global collaboration to map and characterize all cells in a healthy human body."
Getting More Science from Less Data
Images are everywhere. Smart phones and sensors have produced a treasure trove of pictures, many tagged with pertinent information identifying content. Using this vast database of cross-referenced images, convolutional neural networks and other machine learning methods have revolutionized our ability to quickly identify natural images that look like ones previously seen and catalogued.
These methods "learn" by tuning a stunningly large set of hidden internal parameters, guided by millions of tagged images, and requiring large amounts of supercomputer time. But what if you don't have so many tagged images? In many fields, such a database is an unachievable luxury. Biologists record cell images and painstakingly outline the borders and structure by hand: it's not unusual for one person to spend weeks coming up with a single fully three-dimensional image. Materials scientists use tomographic reconstruction to peer inside rocks and materials, and then roll up their sleeves to label different regions, identifying cracks, fractures, and voids by hand. Contrasts between different yet important structures are often very small and "noise" in the data can mask features and confuse the best of algorithms (and humans).
These precious hand-curated images are nowhere near enough for traditional machine learning methods. To meet this challenge, mathematicians at CAMERA attacked the problem of machine learning from very limited amounts of data. Trying to do "more with less," their goal was to figure out how to build an efficient set of mathematical "operators" that could greatly reduce the number of parameters. These mathematical operators might naturally incorporate key constraints to help in identification, such as by including requirements on scientifically plausible shapes and patterns.
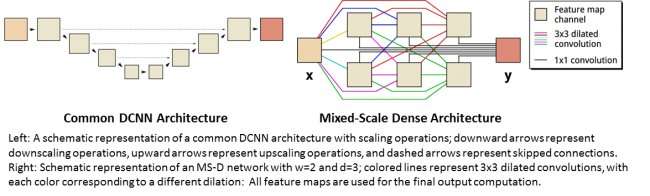
Mixed-Scale Dense Convolution Neural Networks
Many applications of machine learning to imaging problems use deep convolutional neural networks (DCNNs), in which the input image and intermediate images are convolved in a large number of successive layers, allowing the network to learn highly nonlinear features. To achieve accurate results for difficult image processing problems, DCNNs typically rely on combinations of additional operations and connections including, for example, downscaling and upscaling operations to capture features at various image scales. To train deeper and more powerful networks, additional layer types and connections are often required. Finally, DCNNs typically use a large number of intermediate images and trainable parameters, often more than 100 million, to achieve results for difficult problems.
Instead, the new "Mixed-Scale Dense" network architecture avoids many of these complications and calculates dilated convolutions as a substitute to scaling operations to capture features at various spatial ranges, employing multiple scales within a single layer, and densely connecting all intermediate images. The new algorithm achieves accurate results with few intermediate images and parameters, eliminating both the need to tune hyperparameters and additional layers or connections to enable training.
Getting high resolution science from low resolution data
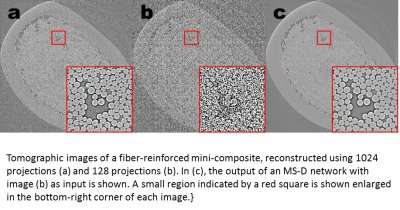
A different challenge is to produce high resolution images from low resolution input. As anyone who has tried to enlarge a small photo and found it only gets worse as it gets bigger, this sounds close to impossible. But a small set of training images processed with a Mixed-Scale Dense network can provide real headway. As an example, imagine trying to denoise tomographic reconstructions of a fiber-reinforced mini-composite material. In an experiment described in the paper, images were reconstructed using 1,024 acquired X-ray projections to obtain images with relatively low amounts of noise. Noisy images of the same object were then obtained by reconstructing using 128 projections. Training inputs were noisy images, with corresponding noiseless images used as target output during training. The trained network was then able to effectively take noisy input data and reconstruct higher resolution images.
New Applications
Pelt and Sethian are taking their approach to a host of new areas, such as fast real-time analysis of images coming out of synchrotron light sources and reconstruction problems in biological reconstruction such as for cells and brain mapping.
"These new approaches are really exciting, since they will enable the application of machine learning to a much greater variety of imaging problems than currently possible," Pelt said. "By reducing the amount of required training images and increasing the size of images that can be processed, the new architecture can be used to answer important questions in many research fields."
More information: Daniël M. Pelt et al. A mixed-scale dense convolutional neural network for image analysis, Proceedings of the National Academy of Sciences (2017). DOI: 10.1073/pnas.1715832114