Computer scientists develop program to find 'low-frequency' variants in sequence data
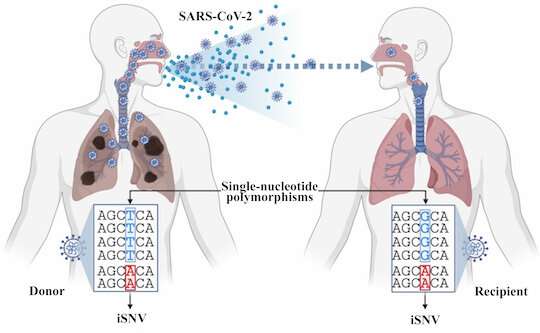
Details about variants hiding in the deluge of genetic SARS-CoV-2 sequences would be good to know, if only researchers can get to them.
A new program developed at Rice University's George R. Brown School of Engineering will make it possible, at least for "intrahost variants," those that appear in genome data from the same COVID-19-positive person.
A Rice team led by computer scientist Todd Treangen and graduate student Yunxi Li has developed Variabel, which accurately identifies "low-frequency variants" of the virus that causes COVID-19.
Finding these clues could be key to identifying potentially devastating variants before they have a chance to spread, Treangen said.
The data is freely available, but there's a lot of it. The research makes low-frequency variant mining available for an estimated half-million SARS-CoV-2 genomes gathered by Oxford Nanopore Technologies (ONT), which offers an affordable platform for rapid sequencing of single, long molecules of DNA or RNA.
"Variabel directly enables the use of affordable nanopore sequencing technology for the identification of within-host variation after viral infection," said Treangen, whose work has focused on infectious disease monitoring since long before the COVID-19 pandemic.
The lab had similar success in testing Variabel on sequence data from patients infected with Ebola and norovirus.
The open-source program, detailed in Nature Communications, is available for download at https://gitlab.com/treangenlab/variabel.
The researchers claim the key to Variabel is its ability to distinguish true variants from sequencing errors in the ONT process.
To validate Variabel, they compared data taken over time from single positive patients as well as sequences from cross-patient datasets, produced by ONT and another sequencing technique, Illumina. Over time, a single patient can host as many as a billion copies of a virus.
By comparing results before and after applying Variabel to the data, they found the program was able to correct the great majority of sequencing errors.
"Variabel opens the door to portable, affordable and rapid characterization of within-host variation, which ultimately could aid in the discovery of future mutations specific to variants of concern," said Treangen, whose lab, along with Rice's Ken Kennedy Institute, hosted a March 11 symposium to discuss scientific advances spurred by the pandemic.
Co-authors of the paper are Rice undergraduate Joshua Kearney and software engineer Bryce Kille, and Baylor College of Medicine postdoctoral associate Medhat Mahmoud and Fritz Sedlazeck, an associate professor at the Human Genome Sequencing Center. Treangen is an assistant professor of computer science.
More information: Yunxi Liu et al, Rescuing low frequency variants within intra-host viral populations directly from Oxford Nanopore sequencing data, Nature Communications (2022). DOI: 10.1038/s41467-022-28852-1