This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
preprint
trusted source
proofread
Researchers develop large language model for medical knowledge

Researchers from EPFL have just released Meditron, the world's best performing open source large language model tailored to the medical field designed to help guide clinical decision-making.
Large language models (LLMs) are deep learning algorithms trained on vast amounts of text to learn billions of mathematical relationships between words (also called "parameters"). They are familiar to most of us as the algorithmic basis for chatbots like OpenAI's ChatGPT and PaLM, used for Google's Bard. Today's largest models have hundreds of billions of parameters, also costing in the billions of dollars to train.
While massive-scale generalist models like ChatGPT can help users with a range of tasks from emails to poetry, focusing on a specific domain of knowledge can allow the models to be smaller and more accessible. For instance, LLMs that are carefully trained on high-quality medical knowledge can potentially democratize access to evidence-based information to help guide clinical decision-making.
Many efforts have already been made to harness and improve LLMs' medical knowledge and reasoning capabilities but, to date, the resulting AI is either closed source (e.g., MedPaLM and GPT-4) or limited in scale, at around 13-billion parameters, which restricts their access or ability.
Seeking to improve access and representation, researchers in EPFL's School of Computer and Communication Sciences have developed MEDITRON 7B and 70B, a pair of open-source LLMs with 7 and 70 billion parameters respectively, adapted to the medical domain, and described in their article posted to the preprint server arXiv, "MEDITRON-70B: Scaling Medical Pretraining for Large Language Models."
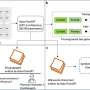
Building on the open-access Llama-2 model released by Meta, with continual input from clinicians and biologists, MEDITRON was trained on carefully curated, high-quality medical data sources. This included peer-reviewed medical literature from open-access repositories like PubMed and a unique set of diverse clinical practice guidelines, covering multiple countries, regions, hospitals, and international organizations.
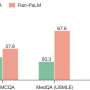
"After developing MEDITRON we evaluated it on four major medical benchmarks showing that its performance exceeds all other open-source models available, as well as the closed GPT-3.5 and Med-PaLM models. MEDITRON-70B is even within 5% of GPT-4 and 10% of Med-PaLM-2, the two best performing, but closed, models currently tailored to medical knowledge," said Zeming Chen, lead author and a doctoral candidate in the Natural Language Processing Lab (NLP) of Professor Antoine Bosselut, the principal investigator of the project.
In a world where many people are suspicious, or even fearful, of the rapid advance of artificial intelligence, Professor Martin Jaggi, head of the Machine Learning and Optimization Laboratory (MLO), emphasizes the importance of EPFL's MEDITRON being open-source, including the code for curating the medical pretraining corpus and the model weights.
"There's transparency in how MEDITRON was trained and what data was used. We want researchers to stress test our model and make it more reliable and robust with their improvements, building on the safety of the tool in the long and necessary process of real-world validation. None of this is available with the closed models developed by big tech," he explained.
Professor Mary-Anne Hartley, a medical doctor and head of the Laboratory for intelligent Global Health Technologies, hosted jointly in the MLO and Yale School of Medicine, is leading the medical aspects of the study. "We designed MEDITRON from the outset with safety in mind. What is unique is that it encodes medical knowledge from transparent sources of high-quality evidence. Now comes the important work of ensuring that the model is able to deliver this information appropriately and safely."
One of these sources of high-quality evidence is the International Committee of the Red Cross clinical practice guidelines.
"It is not often that new health tools are sensitive to the needs of humanitarian contexts," says Dr. Javier Elkin, who heads the Digital Health Program at the International Committee for the Red Cross. "The ICRC is a key custodian of humanitarian principles and we are excited to collaborate with this EPFL initiative that allows us to incorporate our guidelines into the technology."
In early December, a joint workshop in Geneva will explore the potential—as well as the limitations and risks—of this kind of technology, with a special session on MEDITRON from the authors.
"We developed MEDITRON because access to medical knowledge should be a universal right," concluded Bosselut. "We hope that it will prove to be a useful starting point for researchers looking to safely adapt and validate this technology in their practice."
More information: Zeming Chen et al, MEDITRON-70B: Scaling Medical Pretraining for Large Language Models, arXiv (2023). DOI: 10.48550/arxiv.2311.16079