Enzymes could make it cheaper to recycle waste polyester textiles and bottles than making them from petroleum
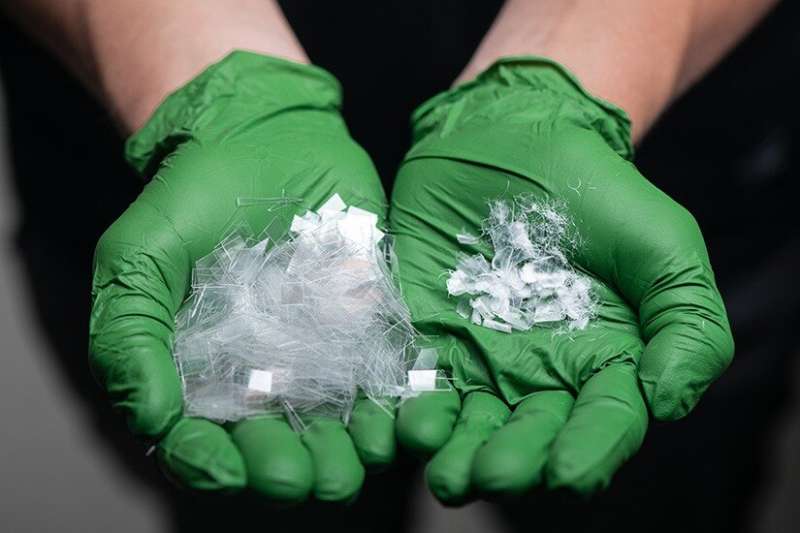

What do a T-shirt, a rug, and a soda bottle have in common? Many are made from polyethylene terephthalate (PET), a ubiquitous plastic that revolutionized the materials industry after it was patented in the 1940s.
Created from petroleum refining, PET is a material known for its durability and versatility. It is easily molded into airtight containers, woven into durable carpets, or spun into polyester clothing.
"The reality is that most PET products—especially PET clothing and carpeting—are not recycled today using conventional recycling technologies," explained Gregg Beckham, senior research fellow at the National Renewable Energy Laboratory (NREL) and CEO of the U.S. Department of Energy BOTTLE Consortium. "The research community is developing promising alternatives, including enzymes designed to depolymerize PET, but even these options have tended to lean on energy-intensive and costly preprocessing steps to be effective."
As a result, most PET produced today eventually finds its way into landfills or the environment—even PET products that actually make it into the recycling bin.
Still, Beckham said that narrative is changing quickly. Advanced methods in machine learning and synthetic biology have given scientists an unprecedented look into the fundamental biology of PET-deconstructing enzymes. As detailed in an article published in Nature Communications, Beckham and colleagues at University of Portsmouth and Montana State University used these methods to discover new enzyme variants that show promise at deconstructing even the toughest PET without extra preprocessing.
Not only could that mean we are on the cusp of enzymatic recycling for all forms of PET, including carpeting and clothing—it means it could soon be cheaper to recycle PET than making it from scratch with petroleum.
Extraordinary enzymes hidden in the dirt
The concept of enzymatic recycling of PET has been known since 2005, but it surged onto the world stage in 2016 after Japanese scientists made a surprising discovery. Buried in the dirt outside a recycling facility in Japan, a bacterium they dubbed Ideonella sakaiensis was quietly secreting enzymes that were deconstructing old plastic beverage bottles scattered about.

Nature had provided an elegant solution for cleaving the chemical bonds of PET. Somehow, nature was demonstrating how to turn PET bottles back to the basic components from which they were made: terephthalic acid and ethylene glycol.
A flurry of research followed. Scientists sought to enhance the enzymes for use in industrial technologies equipped to deal with the millions of tons of PET produced every year. If improved, they hypothesized, an enzymatic recycling platform could overhaul today's underperforming recycling systems, reduce energy and greenhouse gas emissions, and foster a circular economy for all PET products—even carpeting and fabrics not recyclable with conventional technologies.
"The scientific literature lit up with new papers from all over the world as researchers realized the potential of utilizing enzymes to break down plastics," said John McGeehan, a scientist who led the contribution from the team at the University of Portsmouth (UoP) in the United Kingdom. "Experts from fields as diverse as pharma and biofuels were able to repurpose decades of research experience engineering enzymes."
Indeed, Beckham, McGeehan, and colleagues were themselves core players in the global effort to improve enzymatic PET recycling. In a series of frequently cited papers published between 2018 and 2021, the team characterized the enzymes, improved their efficiency sixfold, and analyzed the environmental and economic impact of industrial-scale PET recycling. A corresponding 2022 paper assessed life cycle impacts for a future enzymatic PET recycling system.
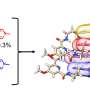
Those studies earned measurable results. By treating PET for 48 hours in a bioreactor, they showed it was possible to transform nearly 98% of the plastic back into terephthalic acid and ethylene glycol—high-quality recycled building blocks for making new PET bottles or even advanced plastics designed to be more easily recyclable.
"It was an incredible experience to be in a field that was expanding so quickly," McGeehan added. "We are reaching a point where collaborative science has huge potential to accelerate the development and rollout of enzyme-based solutions at scale."
Despite rapid advances, though, a key barrier to industrial-scale enzymatic recycling loomed.
The enzymes were only effective on a small percentage of PET products—those made from "amorphous" PET. They struggled to deconstruct the highly common, durable "crystalline" varieties of PET without first softening them with high heat and extra energy. Crystalline PET comprises nearly 90% of all PET produced, including polyester fibers in clothing and parts of single-use beverage bottles.
What the scientists needed were enzymes more adept at breaking down crystalline PET. If only they could find other plastic-eating enzymes hidden somewhere in the dirt.
'Digging' for new enzymes with machine learning
Fortunately, the team did not need a shovel to dig for new enzyme varieties. Advances in bioinformatics and machine learning had already made it possible to prospect vast databases of existing enzyme sequences for varieties active on crystalline PET.
"Traditional approaches to map out new plastic-eating enzymes from databases can be ineffective, since enzymes that are very similar in their chemical makeup may not necessarily retain the plastic deconstruction activity," said NREL computational scientist Japheth Gado.
To get around that challenge, Gado built a statistical model to learn the biological rules of known plastic-deconstructing enzymes. The model assigned probabilities to the unique composition of enzymes studied to date. Gado also built a companion machine-learning model to predict the heat tolerance of enzymes, important for industrial applications.
Together, the two computational models let Gado and colleagues project into the unknown. In less than an hour, they screened over 250 million proteins to create a short list of promising candidates. Further testing confirmed that 36 were able to deconstruct PET, and 24 were previously undescribed in scientific literature.
Importantly, several were even better at breaking down crystalline PET than amorphous PET.
"Not only are these new enzymes genetically diverse," Gado explained. "They are structurally diverse with varying geometries of the active sites."
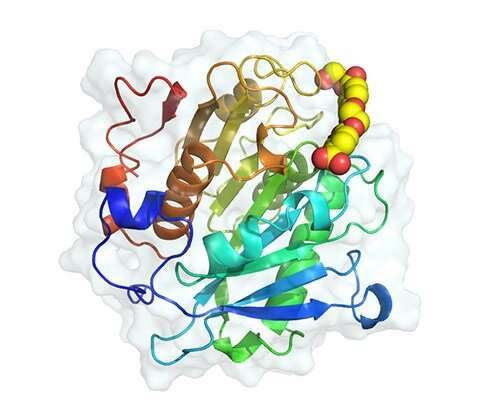
Gado can speak confidently about the structure of the 24 novel enzymes because he has seen what they look like—at least in 3D renderings provided by researchers at DeepMind, a subsidiary of Alphabet. Known for mapping the "entire protein universe" with its deep-learning tool AlphaFold, DeepMind characterized the enzymes so the team could compare the enzymes side by side and note their differences.
All were capable of deconstructing PET, but several looked strikingly different from one another. According to Gado, the renderings from DeepMind provide invaluable clues into how plastic-deconstructing enzymes act on PET.
"State-of-the-art AI methods help us find patterns in the enzyme data that will provide enhanced understanding on what makes a good plastic-eating enzyme," Gado added. "This will enable us to improve the enzymes with protein engineering and to find other enzymes in nature that are similar in terms of performance."
It was another advance from an already prolific research team and one step closer to realizing PET recycling at scale.
Cheaper and greener: Analysis quantifies advantages of enzymatic PET recycling
Cleaning, shredding, and heating—steps needed to prepare PET for deconstruction—are among the most important sustainability drivers of an industrial-scale enzymatic recycling facility, according to Beckham.
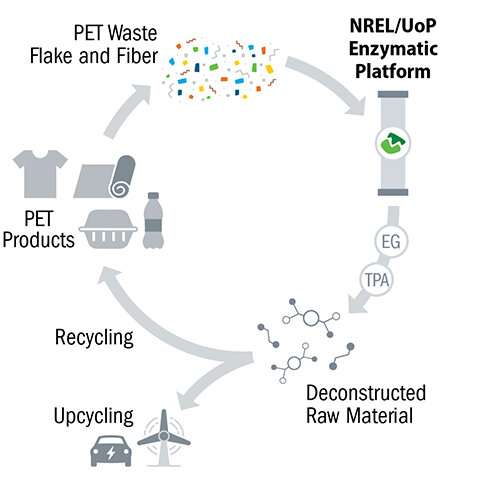
"Minimizing these preprocessing steps is critical for making enzymatic recycling cost competitive with creating PET resin from petroleum," he explained.
In follow-up experiments, the team noted that some of the enzymes flagged by their machine learning methods were equally effective at breaking down crystalline and amorphous PET. Those enzymes simply did not need preprocessing to help soften the bonds of the plastic.
"By cutting out preprocessing, the technology could enable industrial-scale PET recycling that is actually cheaper than virgin PET production using petroleum," Beckham added. "Even better, it can lower associated energy and greenhouse gas emissions."
In an earlier article published in Joule in 2021, the team had already quantified the economic and environmental advantages of using enzymes active on crystalline PET. At an industrial-scale facility, doing so could reduce supply chain energy requirements by 45% and life-cycle greenhouse gas emissions by 38% compared to systems using preprocessing.
The economic advantages are equally as impressive. When deconstructing waste PET carpeting and clothing—unrecyclable with conventional technologies—they could also produce terephthalic acid for less than $1 per kilogram. Petroleum-derived terephthalic acid has historically sold for between $1 and $1.50 per kilogram.
"Our enzymatic platform creates an economic incentive for cleaning up our oceans," said Erika Erickson, a former NREL postdoctoral researcher who conducted much of the experimental work behind the studies. "At such a price point, PET pollution can be affordably recycled into new PET products or find a new purpose in wind turbine blades or carbon fiber bumpers."
Post-consumer PET products, often a source of pollution today, can be transformed into valuable resources to support a more environmentally sustainable plastics economy.
It is not difficult to imagine how this might flip the narrative on the plastic: PET so cheap to recycle that economics favor tossing it in the recycling bin over the trash can. A T-shirt, a rug, a soda bottle—all go in and begin their circular journey as building blocks to create a cleaner, greener world.
More information: Erika Erickson et al, Sourcing thermotolerant poly(ethylene terephthalate) hydrolase scaffolds from natural diversity, Nature Communications (2022). DOI: 10.1038/s41467-022-35237-x